class: center, middle, inverse, title-slide # Spatial data visualization ## Statistical Computing & Programming ### Shawn Santo --- ## Supplementary materials Full video lecture available in Zoom Cloud Recordings Additional resources - Simple Features for R [vignettes](https://r-spatial.github.io/sf/) - [CRS in R](https://www.nceas.ucsb.edu/~frazier/RSpatialGuides/OverviewCoordinateReferenceSystems.pdf) by Melanie Frazier - [Leaflet for R](https://rstudio.github.io/leaflet/) --- ## Data Data for today is available at - Package `sf` - NC counties, births, sudden infant deaths ``` nc <- st_read(system.file("shape/nc.shp", package = "sf")) ``` - [NC OneMap](https://www.nconemap.gov) - lots of spatial data on all things NC - [NC PMTW Streams 2020](https://www.nconemap.gov/datasets/0cc135e6c6244c9e9646b45ee3cb6c1e_0) - [NC game lands](https://www.nconemap.gov/datasets/ncwrc::game-lands-general) - [London cholera data](http://blog.rtwilson.com/john-snows-cholera-data-in-more-formats/) --- class: inverse, center, middle # A little history --- ## Spatial data analysis then John Snow's map that changed the world... <br/> <center> <img src="images/cholera_map.jpg"> </center> Backstory: read more [here](https://www.theguardian.com/news/datablog/2013/mar/15/john-snow-cholera-map) --- ## Spatial data analysis now Spatial data analysis and visualization was a critical component in Operation Neptune Spear. <center> <img src="images/bin_laden_compound.jpeg" width=600 height=400> </center> Backstory: read more [here](https://www.theatlantic.com/politics/archive/2011/05/the-little-known-agency-that-helped-kill-bin-laden/238454/) --- class: inverse, center, middle # Introduction --- ## Spatial data is different Our **typical tidy data frame**: .tiny[ ``` #> # A tibble: 336,776 x 19 #> year month day dep_time sched_dep_time dep_delay arr_time sched_arr_time #> <int> <int> <int> <int> <int> <dbl> <int> <int> #> 1 2013 1 1 517 515 2 830 819 #> 2 2013 1 1 533 529 4 850 830 #> 3 2013 1 1 542 540 2 923 850 #> 4 2013 1 1 544 545 -1 1004 1022 #> 5 2013 1 1 554 600 -6 812 837 #> 6 2013 1 1 554 558 -4 740 728 #> 7 2013 1 1 555 600 -5 913 854 #> 8 2013 1 1 557 600 -3 709 723 #> 9 2013 1 1 557 600 -3 838 846 #> 10 2013 1 1 558 600 -2 753 745 #> # … with 336,766 more rows, and 11 more variables: arr_delay <dbl>, #> # carrier <chr>, flight <int>, tailnum <chr>, origin <chr>, dest <chr>, #> # air_time <dbl>, distance <dbl>, hour <dbl>, minute <dbl>, time_hour <dttm> ``` ] --- ## Spatial data is different A **simple features object**: .tiny[ ``` #> Simple feature collection with 100 features and 5 fields #> geometry type: MULTIPOLYGON #> dimension: XY #> bbox: xmin: -84.32385 ymin: 33.88199 xmax: -75.45698 ymax: 36.58965 #> geographic CRS: NAD27 #> First 10 features: #> AREA PERIMETER CNTY_ CNTY_ID NAME geometry #> 1 0.114 1.442 1825 1825 Ashe MULTIPOLYGON (((-81.47276 3... #> 2 0.061 1.231 1827 1827 Alleghany MULTIPOLYGON (((-81.23989 3... #> 3 0.143 1.630 1828 1828 Surry MULTIPOLYGON (((-80.45634 3... #> 4 0.070 2.968 1831 1831 Currituck MULTIPOLYGON (((-76.00897 3... #> 5 0.153 2.206 1832 1832 Northampton MULTIPOLYGON (((-77.21767 3... #> 6 0.097 1.670 1833 1833 Hertford MULTIPOLYGON (((-76.74506 3... #> 7 0.062 1.547 1834 1834 Camden MULTIPOLYGON (((-76.00897 3... #> 8 0.091 1.284 1835 1835 Gates MULTIPOLYGON (((-76.56251 3... #> 9 0.118 1.421 1836 1836 Warren MULTIPOLYGON (((-78.30876 3... #> 10 0.124 1.428 1837 1837 Stokes MULTIPOLYGON (((-80.02567 3... ``` ] --- Another **simple features** object: .tiny[ ``` #> Simple feature collection with 1858 features and 2 fields #> geometry type: MULTILINESTRING #> dimension: XYZ #> bbox: xmin: -9383722 ymin: 4162088 xmax: -8947168 ymax: 4381832 #> z_range: zmin: 0 zmax: 0 #> projected CRS: WGS 84 / Pseudo-Mercator #> First 10 features: #> Displ_Name FIRST_WRC_ #> 1 Big Sandy Creek Delayed Harvest Trout Waters #> 2 Helton Creek Delayed Harvest Trout Waters #> 3 Little River Delayed Harvest Trout Waters #> 4 Cranberry Creek Hatchery Supported Trout Waters #> 5 Big Pine Creek Hatchery Supported Trout Waters #> 6 Bledsoe Creek Hatchery Supported Trout Waters #> 7 Brush Creek Hatchery Supported Trout Waters #> 8 <NA> Hatchery Supported Trout Waters #> 9 (Big) Glade Creek Hatchery Supported Trout Waters #> 10 Little River Hatchery Supported Trout Waters #> geometry #> 1 MULTILINESTRING Z ((-901773... #> 2 MULTILINESTRING Z ((-907349... #> 3 MULTILINESTRING Z ((-903290... #> 4 MULTILINESTRING Z ((-904529... #> 5 MULTILINESTRING Z ((-901483... #> 6 MULTILINESTRING Z ((-903607... #> 7 MULTILINESTRING Z ((-901813... #> 8 MULTILINESTRING Z ((-902513... #> 9 MULTILINESTRING Z ((-902642... #> 10 MULTILINESTRING Z ((-903124... ``` ] --- Yet another **simple features** object: .tiny[ ``` #> Simple feature collection with 94 features and 2 fields #> geometry type: MULTIPOLYGON #> dimension: XY #> bbox: xmin: 127456.7 ymin: 26544.91 xmax: 923528.7 ymax: 318097.4 #> projected CRS: NAD83 / North Carolina #> First 10 features: #> GML_HAB SUM_ACRES geometry #> 1 Alcoa 11109.5589 MULTIPOLYGON (((512096.2 18... #> 2 Alligator River 24439.0891 MULTIPOLYGON (((869633.1 24... #> 3 Angola Bay 34067.3821 MULTIPOLYGON (((713079.4 11... #> 4 Bachelor Bay 2786.2577 MULTIPOLYGON (((813742.2 23... #> 5 Bertie County 3881.4660 MULTIPOLYGON (((797133.8 24... #> 6 Bladen Lakes State Forest 33671.8439 MULTIPOLYGON (((658970.6 95... #> 7 Brinkleyville 3189.7974 MULTIPOLYGON (((714741 2769... #> 8 Buckhorn 491.3477 MULTIPOLYGON (((589723.7 25... #> 9 Buckridge 17965.7187 MULTIPOLYGON (((871137.4 21... #> 10 Buffalo Cove 6630.9453 MULTIPOLYGON (((381445.9 26... ``` ] --- ## Spatial data plotting needs care <img src="lec_11_files/figure-html/unnamed-chunk-6-1.png" width="100%" style="display: block; margin: auto;" /> ``` #> null device #> 1 ``` --- <img src="lec_11_files/figure-html/unnamed-chunk-8-1.png" width="100%" style="display: block; margin: auto;" /> --- <img src="lec_11_files/figure-html/unnamed-chunk-9-1.png" width="100%" style="display: block; margin: auto;" /> --- class: center, middle ## Can we combine the two plots? --- ## What happened? <img src="lec_11_files/figure-html/unnamed-chunk-10-1.png" width="100%" style="display: block; margin: auto;" /> --- class: center, middle ## We can combine them, but more care is needed --- <img src="lec_11_files/figure-html/unnamed-chunk-11-1.png" width="100%" style="display: block; margin: auto;" /> --- ## Spatial data challenges 1. Different data types exist. 2. Special attention must be given to the coordinate reference system (CRS). 3. Manipulating spatial data objects is similar but not identical to manipulating data frame objects. --- class: inverse, center, middle # Spatial data and R --- ## Analysis of spatial data in R .pull-left[ <br/> - Package `raster` contains classes and tools for handling spatial raster data. <br/><br/> - Package `sf` combines the functionality of `sp`, `rgdal`, and `rgeos` into a single package based on tidy simple features. ] .pull-right[ 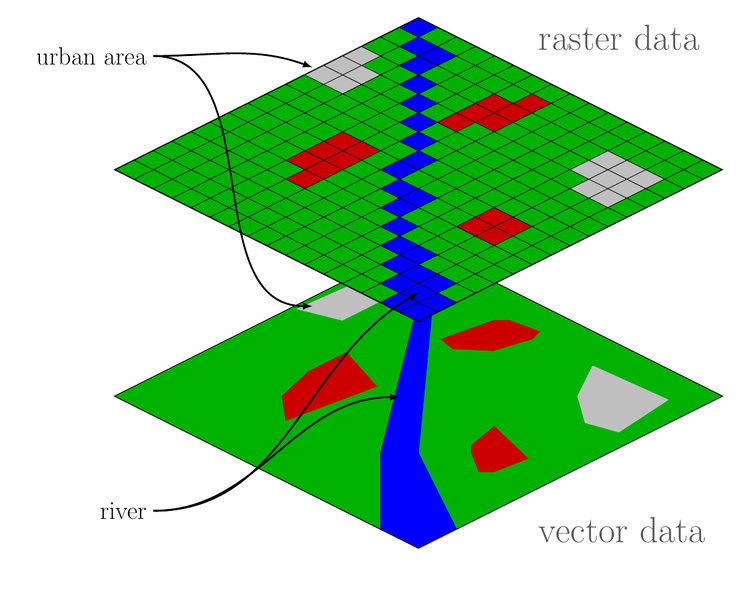 ] <br/> Whether or not you use vector or raster data depends on the type of problem and the data source. Our focus will be on vector data and package `sf`. *Source:* https://commons.wikimedia.org/wiki/File:Raster_vector_tikz.png --- ## Installing package `sf` From https://r-spatial.github.io/sf/index.html **Windows** Installing `sf` from source works under windows when Rtools is installed. This downloads the system requirements from rwinlib. **MacOS** ```bash brew install pkg-config brew install gdal ``` Once gdal is installed, you will be able to install sf package from source in R. **Linux** For Unix-alikes, GDAL (>= 2.0.1), GEOS (>= 3.4.0) and Proj.4 (>= 4.8.0) are required. --- ## Features and simple features - A **feature** is a thing or object in the real world: a house, a city, a park, a forest, etc. <br/><br/> - A **simple feature** as defined by OpenGIS Abstract specification is to have both spatial and non-spatial attributes. Spatial attributes are geometry valued, and simple features are based on 2D geometry with linear interpolation between vertices. <br/><br/> .tiny[ ```r Simple feature collection with 1858 features and 2 fields geometry type: MULTILINESTRING dimension: XYZ bbox: xmin: -9383722 ymin: 4162088 xmax: -8947168 ymax: 4381832 z_range: zmin: 0 zmax: 0 projected CRS: WGS 84 / Pseudo-Mercator First 10 features: Displ_Name FIRST_WRC_ geometry *1 Big Sandy Creek Delayed Harvest Trout Waters MULTILINESTRING Z ((-901773... 2 Helton Creek Delayed Harvest Trout Waters MULTILINESTRING Z ((-907349... 3 Little River Delayed Harvest Trout Waters MULTILINESTRING Z ((-903290... 4 Cranberry Creek Hatchery Supported Trout Waters MULTILINESTRING Z ((-904529... 5 Big Pine Creek Hatchery Supported Trout Waters MULTILINESTRING Z ((-901483... 6 Bledsoe Creek Hatchery Supported Trout Waters MULTILINESTRING Z ((-903607... 7 Brush Creek Hatchery Supported Trout Waters MULTILINESTRING Z ((-901813... 8 <NA> Hatchery Supported Trout Waters MULTILINESTRING Z ((-902513... 9 (Big) Glade Creek Hatchery Supported Trout Waters MULTILINESTRING Z ((-902642... 10 Little River Hatchery Supported Trout Waters MULTILINESTRING Z ((-903124... ``` ] --- ## Simple features examples <img src="lec_11_files/figure-html/unnamed-chunk-14-1.png" width="100%" style="display: block; margin: auto;" /> ``` #> null device #> 1 ``` --- ## `sf` objects .tiny[ ```r nc <- st_read(system.file("shape/nc.shp", package = "sf"), quiet = TRUE) nc ``` ``` #> Simple feature collection with 100 features and 14 fields #> geometry type: MULTIPOLYGON #> dimension: XY #> bbox: xmin: -84.32385 ymin: 33.88199 xmax: -75.45698 ymax: 36.58965 #> geographic CRS: NAD27 #> First 10 features: #> AREA PERIMETER CNTY_ CNTY_ID NAME FIPS FIPSNO CRESS_ID BIR74 SID74 #> 1 0.114 1.442 1825 1825 Ashe 37009 37009 5 1091 1 #> 2 0.061 1.231 1827 1827 Alleghany 37005 37005 3 487 0 #> 3 0.143 1.630 1828 1828 Surry 37171 37171 86 3188 5 #> 4 0.070 2.968 1831 1831 Currituck 37053 37053 27 508 1 #> 5 0.153 2.206 1832 1832 Northampton 37131 37131 66 1421 9 #> 6 0.097 1.670 1833 1833 Hertford 37091 37091 46 1452 7 #> 7 0.062 1.547 1834 1834 Camden 37029 37029 15 286 0 #> 8 0.091 1.284 1835 1835 Gates 37073 37073 37 420 0 #> 9 0.118 1.421 1836 1836 Warren 37185 37185 93 968 4 #> 10 0.124 1.428 1837 1837 Stokes 37169 37169 85 1612 1 #> NWBIR74 BIR79 SID79 NWBIR79 geometry #> 1 10 1364 0 19 MULTIPOLYGON (((-81.47276 3... #> 2 10 542 3 12 MULTIPOLYGON (((-81.23989 3... #> 3 208 3616 6 260 MULTIPOLYGON (((-80.45634 3... #> 4 123 830 2 145 MULTIPOLYGON (((-76.00897 3... #> 5 1066 1606 3 1197 MULTIPOLYGON (((-77.21767 3... #> 6 954 1838 5 1237 MULTIPOLYGON (((-76.74506 3... #> 7 115 350 2 139 MULTIPOLYGON (((-76.00897 3... #> 8 254 594 2 371 MULTIPOLYGON (((-76.56251 3... #> 9 748 1190 2 844 MULTIPOLYGON (((-78.30876 3... #> 10 160 2038 5 176 MULTIPOLYGON (((-80.02567 3... ``` ] --- ## Class and other attributes: `sf` ```r class(nc) ``` ``` #> [1] "sf" "data.frame" ``` ```r names(attributes(nc)) ``` ``` #> [1] "names" "row.names" "class" "sf_column" "agr" ``` --- ## `sfc` objects ```r nc_polygons <- st_geometry(nc) nc_polygons ``` ``` #> Geometry set for 100 features #> geometry type: MULTIPOLYGON #> dimension: XY #> bbox: xmin: -84.32385 ymin: 33.88199 xmax: -75.45698 ymax: 36.58965 #> geographic CRS: NAD27 #> First 5 geometries: ``` ``` #> MULTIPOLYGON (((-81.47276 36.23436, -81.54084 3... ``` ``` #> MULTIPOLYGON (((-81.23989 36.36536, -81.24069 3... ``` ``` #> MULTIPOLYGON (((-80.45634 36.24256, -80.47639 3... ``` ``` #> MULTIPOLYGON (((-76.00897 36.3196, -76.01735 36... ``` ``` #> MULTIPOLYGON (((-77.21767 36.24098, -77.23461 3... ``` --- ## Class and other attributes: `sfc` ```r class(nc_polygons) ``` ``` #> [1] "sfc_MULTIPOLYGON" "sfc" ``` ```r names(attributes(nc_polygons)) ``` ``` #> [1] "n_empty" "crs" "class" "precision" "bbox" ``` <br/> -- We see that `nc` has a class attribute `sf`, and object `nc_polygons` has a class attribute `sfc`. What methods are available? --- ```r sloop::s3_methods_class("sf") %>% print(n = 20) ``` ``` #> # A tibble: 92 x 4 #> generic class visible source #> <chr> <chr> <lgl> <chr> #> 1 [ sf FALSE registered S3method #> 2 [[<- sf FALSE registered S3method #> 3 $<- sf FALSE registered S3method #> 4 aggregate sf FALSE registered S3method #> 5 anti_join sf FALSE registered S3method #> 6 arrange sf FALSE registered S3method #> 7 as.data.frame sf FALSE registered S3method #> 8 cbind sf FALSE registered S3method #> 9 distinct sf FALSE registered S3method #> 10 dplyr_reconstruct sf FALSE registered S3method #> 11 filter sf FALSE registered S3method #> 12 full_join sf FALSE registered S3method #> 13 gather sf FALSE registered S3method #> 14 group_by sf FALSE registered S3method #> 15 group_split sf FALSE registered S3method #> 16 identify sf FALSE registered S3method #> 17 inner_join sf FALSE registered S3method #> 18 left_join sf FALSE registered S3method #> 19 merge sf FALSE registered S3method #> 20 mutate sf FALSE registered S3method #> # … with 72 more rows ``` --- ```r sloop::s3_methods_class("sfc") %>% print(n = 20) ``` ``` #> # A tibble: 65 x 4 #> generic class visible source #> <chr> <chr> <lgl> <chr> #> 1 [ sfc FALSE registered S3method #> 2 [<- sfc FALSE registered S3method #> 3 as.data.frame sfc FALSE registered S3method #> 4 c sfc FALSE registered S3method #> 5 format sfc FALSE registered S3method #> 6 fortify sfc FALSE registered S3method #> 7 identify sfc FALSE registered S3method #> 8 obj_sum sfc FALSE registered S3method #> 9 Ops sfc FALSE registered S3method #> 10 print sfc FALSE registered S3method #> 11 rep sfc FALSE registered S3method #> 12 scale_type sfc FALSE registered S3method #> 13 st_area sfc FALSE registered S3method #> 14 st_as_binary sfc FALSE registered S3method #> 15 st_as_grob sfc FALSE registered S3method #> 16 st_as_s2 sfc FALSE registered S3method #> 17 st_as_sf sfc FALSE registered S3method #> 18 st_as_text sfc FALSE registered S3method #> 19 st_bbox sfc FALSE registered S3method #> 20 st_boundary sfc FALSE registered S3method #> # … with 45 more rows ``` --- ## Reading and writing spatial data - `st_read()` / `st_write()`, Shapefile, GeoJSON, KML, ... - `st_as_sfc()` - `st_as_text()`, well-known text format - `st_as_binary()`, well-known binary format <br/><br/><br> See https://r-spatial.github.io/sf/articles/sf2.html for the full set of driver availability. --- ## Plotting with `plot()` ```r plot(nc, max.plot = 14) ``` <img src="lec_11_files/figure-html/unnamed-chunk-24-1.png" style="display: block; margin: auto;" /> --- ```r plot(nc["NAME"]) ``` <img src="lec_11_files/figure-html/unnamed-chunk-25-1.png" style="display: block; margin: auto;" /> --- ```r par(oma=c(0,2,0,0)) plot(nc["AREA"], graticule = TRUE, axes = TRUE, las = 1) ``` <img src="lec_11_files/figure-html/unnamed-chunk-26-1.png" style="display: block; margin: auto;" /> --- ## What is happening with `[` and the `sf` object? ```r nc["AREA"] ``` ``` #> Simple feature collection with 100 features and 1 field #> geometry type: MULTIPOLYGON #> dimension: XY #> bbox: xmin: -84.32385 ymin: 33.88199 xmax: -75.45698 ymax: 36.58965 #> geographic CRS: NAD27 #> First 10 features: #> AREA geometry #> 1 0.114 MULTIPOLYGON (((-81.47276 3... #> 2 0.061 MULTIPOLYGON (((-81.23989 3... #> 3 0.143 MULTIPOLYGON (((-80.45634 3... #> 4 0.070 MULTIPOLYGON (((-76.00897 3... #> 5 0.153 MULTIPOLYGON (((-77.21767 3... #> 6 0.097 MULTIPOLYGON (((-76.74506 3... #> 7 0.062 MULTIPOLYGON (((-76.00897 3... #> 8 0.091 MULTIPOLYGON (((-76.56251 3... #> 9 0.118 MULTIPOLYGON (((-78.30876 3... #> 10 0.124 MULTIPOLYGON (((-80.02567 3... ``` --- ```r nc$AREA ``` ``` #> [1] 0.114 0.061 0.143 0.070 0.153 0.097 0.062 0.091 0.118 0.124 0.114 0.153 #> [13] 0.143 0.109 0.072 0.190 0.053 0.199 0.081 0.063 0.044 0.064 0.086 0.128 #> [25] 0.108 0.170 0.111 0.180 0.104 0.077 0.142 0.059 0.131 0.122 0.080 0.118 #> [37] 0.219 0.118 0.155 0.069 0.066 0.145 0.134 0.100 0.099 0.116 0.201 0.180 #> [49] 0.094 0.134 0.168 0.106 0.168 0.207 0.144 0.094 0.203 0.141 0.070 0.065 #> [61] 0.146 0.142 0.154 0.118 0.078 0.125 0.181 0.143 0.091 0.130 0.103 0.095 #> [73] 0.078 0.104 0.098 0.091 0.060 0.131 0.241 0.082 0.120 0.172 0.121 0.163 #> [85] 0.138 0.098 0.167 0.204 0.121 0.051 0.177 0.080 0.195 0.240 0.125 0.225 #> [97] 0.214 0.240 0.042 0.212 ``` --- ```r par(oma=c(0,2,0,0)) plot(nc["AREA"], col = "lightblue", graticule = TRUE, axes = TRUE, las = 1) ``` <img src="lec_11_files/figure-html/unnamed-chunk-29-1.png" style="display: block; margin: auto;" /> --- ```r par(oma=c(0,2,0,0)) plot(st_geometry(nc), graticule = TRUE, axes = TRUE, las = 1) ``` <img src="lec_11_files/figure-html/unnamed-chunk-30-1.png" style="display: block; margin: auto;" /> --- ## Plotting with `ggplot()` ```r ggplot(nc) + * geom_sf() + theme_bw(base_size = 16) ``` <img src="lec_11_files/figure-html/unnamed-chunk-31-1.png" style="display: block; margin: auto;" /> --- ```r ggplot(nc) + * geom_sf(aes(fill = AREA)) + scale_fill_gradient(low = "#fee8c8", high = "#7f0000") + theme_bw(base_size = 16) ``` <img src="lec_11_files/figure-html/unnamed-chunk-32-1.png" style="display: block; margin: auto;" /> --- ## Patchwork works <img src="lec_11_files/figure-html/unnamed-chunk-33-1.png" style="display: block; margin: auto;" /> --- ## Patchwork works Aside: What is wrong with the last set of figures? -- <br/> Code for previous slide: ```r p1 <- ggplot(nc) + geom_sf(aes(fill = SID74)) + scale_fill_gradient(low = "#fff7f3", high = "#49006a") + theme_bw(base_size = 16) p2 <- ggplot(nc) + geom_sf(aes(fill = SID79)) + scale_fill_gradient(low = "#fff7f3", high = "#49006a") + theme_bw(base_size = 16) p1 / p2 ``` --- ## Plotting with `leaflet` ```r library(leaflet) ``` -- ```r # create a color palette function pal <- colorNumeric(palette = "Blues", domain = nc$BIR74) leaflet(nc, width = "100%") %>% addTiles() %>% addPolygons(color = "grey") %>% addPolygons(stroke = FALSE, fillOpacity = 0.8, smoothFactor = 0.2, color = ~pal(BIR74)) %>% addLegend(position = "bottomright", pal = pal, values = ~BIR74, opacity = 0.8) ``` --- ## Plotting with `leaflet` <div id="htmlwidget-052685a893911dd7ccb4" style="width:100%;height:504px;" class="leaflet html-widget"></div> <script type="application/json" data-for="htmlwidget-052685a893911dd7ccb4">{"x":{"options":{"crs":{"crsClass":"L.CRS.EPSG3857","code":null,"proj4def":null,"projectedBounds":null,"options":{}}},"calls":[{"method":"addTiles","args":["//{s}.tile.openstreetmap.org/{z}/{x}/{y}.png",null,null,{"minZoom":0,"maxZoom":18,"tileSize":256,"subdomains":"abc","errorTileUrl":"","tms":false,"noWrap":false,"zoomOffset":0,"zoomReverse":false,"opacity":1,"zIndex":1,"detectRetina":false,"attribution":"© <a href=\"http://openstreetmap.org\">OpenStreetMap<\/a> contributors, <a href=\"http://creativecommons.org/licenses/by-sa/2.0/\">CC-BY-SA<\/a>"}]},{"method":"addPolygons","args":[[[[{"lng":[-81.4727554321289,-81.5408401489258,-81.5619812011719,-81.6330642700195,-81.7410736083984,-81.6982803344727,-81.7027969360352,-81.6699981689453,-81.3452987670898,-81.347541809082,-81.3247756958008,-81.3133239746094,-81.2662353515625,-81.2628402709961,-81.2406921386719,-81.2398910522461,-81.2642440795898,-81.3289947509766,-81.3613739013672,-81.3656921386719,-81.354133605957,-81.3674545288086,-81.4063873291016,-81.4123306274414,-81.431037902832,-81.4528884887695,-81.4727554321289],"lat":[36.2343559265137,36.2725067138672,36.2735939025879,36.3406867980957,36.3917846679688,36.4717788696289,36.5193405151367,36.5896492004395,36.5728645324707,36.537914276123,36.5136795043945,36.4806976318359,36.4372062683105,36.4050407409668,36.3794174194336,36.365364074707,36.3524131774902,36.3635025024414,36.3531608581543,36.3390502929688,36.2997169494629,36.2786979675293,36.2850532531738,36.2672920227051,36.2607192993164,36.2395858764648,36.2343559265137]}]],[[{"lng":[-81.2398910522461,-81.2406921386719,-81.2628402709961,-81.2662353515625,-81.3133239746094,-81.3247756958008,-81.347541809082,-81.3452987670898,-80.9034423828125,-80.9335479736328,-80.9657745361328,-80.9496688842773,-80.9563903808594,-80.9779510498047,-80.9828414916992,-81.0027770996094,-81.0246429443359,-81.0428009033203,-81.0842514038086,-81.0985641479492,-81.1133117675781,-81.1293792724609,-81.1383972167969,-81.1533660888672,-81.1766738891602,-81.2398910522461],"lat":[36.365364074707,36.3794174194336,36.4050407409668,36.4372062683105,36.4806976318359,36.5136795043945,36.537914276123,36.5728645324707,36.5652122497559,36.4983139038086,36.4672203063965,36.4147338867188,36.4037971496582,36.3913764953613,36.3718338012695,36.3666801452637,36.3778343200684,36.4103355407715,36.4299201965332,36.43115234375,36.4228515625,36.4263305664062,36.4176254272461,36.4247398376465,36.4154434204102,36.365364074707]}]],[[{"lng":[-80.4563446044922,-80.4763870239258,-80.5368804931641,-80.5450134277344,-80.5541534423828,-80.5905990600586,-80.6243133544922,-80.6674423217773,-80.6966552734375,-80.7240371704102,-80.7343673706055,-80.7525634765625,-80.7662963867188,-80.7826995849609,-80.874382019043,-80.8708648681641,-80.8889236450195,-80.9245681762695,-80.9563903808594,-80.9496688842773,-80.9657745361328,-80.9335479736328,-80.9034423828125,-80.8381576538086,-80.6110534667969,-80.4353103637695,-80.453010559082,-80.4563446044922],"lat":[36.2425575256348,36.2547264099121,36.2567367553711,36.2766571044922,36.2784309387207,36.2682762145996,36.2730979919434,36.2461013793945,36.259090423584,36.258472442627,36.2647590637207,36.2582969665527,36.2618370056152,36.2485771179199,36.2338829040527,36.3246231079102,36.3544273376465,36.3727531433105,36.4037971496582,36.4147338867188,36.4672203063965,36.4983139038086,36.5652122497559,36.5634384155273,36.5572967529297,36.5510444641113,36.2570877075195,36.2425575256348]}]],[[{"lng":[-76.0089721679688,-76.0173492431641,-76.0328750610352,-76.0439529418945,-76.095085144043,-76.1609268188477,-76.1581497192383,-76.1682891845703,-76.3302536010742,-76.1273956298828,-76.0459594726562,-76.0332107543945,-76.091064453125,-75.97607421875,-75.9697647094727,-76.0016098022461,-75.9512557983398,-75.9281234741211,-75.9245910644531,-75.8000564575195,-75.7988510131836,-75.8551635742188,-75.9137649536133,-75.9575119018555,-75.9419326782227,-76.0089721679688],"lat":[36.3195953369141,36.3377304077148,36.3359756469727,36.3535919189453,36.3489151000977,36.3918991088867,36.4126892089844,36.4270858764648,36.5560569763184,36.5571632385254,36.5569534301758,36.5143737792969,36.5035667419434,36.4362144470215,36.4151191711426,36.4189147949219,36.3654708862305,36.4232444763184,36.3509483337402,36.1128158569336,36.0728187561035,36.1056671142578,36.244800567627,36.2594528198242,36.2943382263184,36.3195953369141]}],[{"lng":[-76.0271682739258,-75.998664855957,-75.9119186401367,-75.9248046875,-75.9772796630859,-75.9762878417969,-76.0271682739258],"lat":[36.5567169189453,36.5566520690918,36.5425300598145,36.4739761352539,36.4780158996582,36.5179252624512,36.5567169189453]}],[{"lng":[-75.901985168457,-75.8781661987305,-75.7731552124023,-75.7831726074219,-75.901985168457],"lat":[36.5561981201172,36.5558738708496,36.2292556762695,36.2251930236816,36.5561981201172]}]],[[{"lng":[-77.2176666259766,-77.2346115112305,-77.2986145019531,-77.2935104370117,-77.3094787597656,-77.3349914550781,-77.389030456543,-77.3786239624023,-77.4134674072266,-77.4188537597656,-77.4541168212891,-77.5380783081055,-77.5574340820312,-77.5719528198242,-77.580078125,-77.559684753418,-77.560417175293,-77.6359710693359,-77.650993347168,-77.6988754272461,-77.7494049072266,-77.9012069702148,-77.8988571166992,-77.7639312744141,-77.3200531005859,-77.1773529052734,-77.1544189453125,-77.0921325683594,-77.075309753418,-77.0832748413086,-77.127326965332,-77.1393203735352,-77.141960144043,-77.2176666259766],"lat":[36.2409820556641,36.214599609375,36.2115287780762,36.1628608703613,36.1627655029297,36.1828498840332,36.2031021118164,36.2400856018066,36.2558174133301,36.281795501709,36.3197555541992,36.3024597167969,36.3042144775391,36.3144950866699,36.3282699584961,36.3759460449219,36.4063568115234,36.4405364990234,36.4725875854492,36.4789581298828,36.4735717773438,36.5098876953125,36.5529441833496,36.5534400939941,36.5539169311523,36.5562858581543,36.526252746582,36.5075187683105,36.4835166931152,36.4700965881348,36.4707107543945,36.4564781188965,36.417064666748,36.2409820556641]}]],[[{"lng":[-76.7450637817383,-76.9806900024414,-76.9947509765625,-77.1300735473633,-77.2176666259766,-77.141960144043,-77.1393203735352,-77.127326965332,-77.0832748413086,-77.075309753418,-77.0921325683594,-77.1544189453125,-77.1773529052734,-76.9241333007812,-76.9084243774414,-76.9457702636719,-76.9536743164062,-76.9435119628906,-76.9240798950195,-76.7413482666016,-76.7074966430664,-76.7450637817383],"lat":[36.2339172363281,36.2302360534668,36.2355804443359,36.2334632873535,36.2409820556641,36.417064666748,36.4564781188965,36.4707107543945,36.4700965881348,36.4835166931152,36.5075187683105,36.526252746582,36.5562858581543,36.5541458129883,36.5042839050293,36.4589614868164,36.4192314147949,36.4017295837402,36.3924446105957,36.3151664733887,36.2661323547363,36.2339172363281]}]],[[{"lng":[-76.0089721679688,-75.9571838378906,-75.9813385009766,-76.1831665039062,-76.1934814453125,-76.216194152832,-76.2385330200195,-76.2612838745117,-76.2741317749023,-76.3033599853516,-76.3213653564453,-76.4059677124023,-76.4983444213867,-76.5635833740234,-76.49755859375,-76.3302536010742,-76.1682891845703,-76.1581497192383,-76.1609268188477,-76.095085144043,-76.0439529418945,-76.0328750610352,-76.0173492431641,-76.0089721679688],"lat":[36.3195953369141,36.1937713623047,36.1697273254395,36.3152389526367,36.3248519897461,36.3278465270996,36.3612289428711,36.3637580871582,36.3814086914062,36.391845703125,36.4096450805664,36.4471588134766,36.5039024353027,36.5552520751953,36.5558128356934,36.5560569763184,36.4270858764648,36.4126892089844,36.3918991088867,36.3489151000977,36.3535919189453,36.3359756469727,36.3377304077148,36.3195953369141]}]],[[{"lng":[-76.5625076293945,-76.6042404174805,-76.6482162475586,-76.6887359619141,-76.7766418457031,-76.9240798950195,-76.9435119628906,-76.9536743164062,-76.9457702636719,-76.9084243774414,-76.9241333007812,-76.921630859375,-76.5635833740234,-76.4983444213867,-76.5024642944336,-76.4603500366211,-76.5625076293945],"lat":[36.3405685424805,36.3149833679199,36.315315246582,36.2945175170898,36.3583297729492,36.3924446105957,36.4017295837402,36.4192314147949,36.4589614868164,36.5042839050293,36.5541458129883,36.5541572570801,36.5552520751953,36.5039024353027,36.4522857666016,36.3738975524902,36.3405685424805]}]],[[{"lng":[-78.3087615966797,-78.2829284667969,-78.3212509155273,-78.0516662597656,-77.8988571166992,-77.9012069702148,-77.9169235229492,-77.9301376342773,-77.9520874023438,-78.0062866210938,-78.058349609375,-78.1096267700195,-78.1347198486328,-78.3087615966797],"lat":[36.2600402832031,36.2918815612793,36.5455322265625,36.5524749755859,36.5529441833496,36.5098876953125,36.5006370544434,36.3529853820801,36.2812309265137,36.1959457397461,36.2113227844238,36.213508605957,36.2365837097168,36.2600402832031]}]],[[{"lng":[-80.0256729125977,-80.453010559082,-80.4353103637695,-80.048095703125,-80.024055480957,-80.0256729125977],"lat":[36.2502326965332,36.2570877075195,36.5510444641113,36.5471343994141,36.5450248718262,36.2502326965332]}]],[[{"lng":[-79.5305099487305,-79.5102996826172,-79.2170639038086,-79.1443252563477,-79.1592712402344,-79.2584991455078,-79.5305099487305],"lat":[36.2461357116699,36.5476570129395,36.5497817993164,36.5460586547852,36.2336692810059,36.2356872558594,36.2461357116699]}]],[[{"lng":[-79.5305099487305,-79.5305786132812,-80.0256729125977,-80.024055480957,-79.7174453735352,-79.5102996826172,-79.5305099487305],"lat":[36.2461357116699,36.236156463623,36.2502326965332,36.5450248718262,36.5478897094727,36.5476570129395,36.2461357116699]}]],[[{"lng":[-78.7491226196289,-78.7884140014648,-78.8040542602539,-78.8103561401367,-78.8068008422852,-78.7966995239258,-78.7373886108398,-78.4588088989258,-78.463752746582,-78.5025024414062,-78.51708984375,-78.5147171020508,-78.4925231933594,-78.5458526611328,-78.5480270385742,-78.6955718994141,-78.7491226196289],"lat":[36.063591003418,36.062183380127,36.080940246582,36.114574432373,36.231575012207,36.5435333251953,36.5460739135742,36.5414810180664,36.5238571166992,36.5043907165527,36.461483001709,36.1752243041992,36.1735877990723,36.0680885314941,36.0141258239746,36.0666465759277,36.063591003418]}]],[[{"lng":[-78.8068008422852,-78.9510803222656,-79.1592712402344,-79.1443252563477,-78.7966995239258,-78.8068008422852],"lat":[36.231575012207,36.2338371276855,36.2336692810059,36.5460586547852,36.5435333251953,36.231575012207]}]],[[{"lng":[-78.4925231933594,-78.5147171020508,-78.51708984375,-78.5025024414062,-78.463752746582,-78.4588088989258,-78.3212509155273,-78.2829284667969,-78.3087615966797,-78.3460464477539,-78.3808517456055,-78.4169540405273,-78.4925231933594],"lat":[36.1735877990723,36.1752243041992,36.461483001709,36.5043907165527,36.5238571166992,36.5414810180664,36.5455322265625,36.2918815612793,36.2600402832031,36.2251815795898,36.2247505187988,36.1621742248535,36.1735877990723]}]],[[{"lng":[-77.3322067260742,-77.4053115844727,-77.4257431030273,-77.4382171630859,-77.4639739990234,-77.5251312255859,-77.5358581542969,-77.5366897583008,-77.5478820800781,-77.6062545776367,-77.6423568725586,-77.6856231689453,-77.7005081176758,-77.7201843261719,-77.7471694946289,-77.8013687133789,-77.8113021850586,-77.8868408203125,-77.917839050293,-77.9224014282227,-77.9392318725586,-77.9558639526367,-77.9733963012695,-77.9851150512695,-78.0062866210938,-77.9520874023438,-77.9301376342773,-77.9169235229492,-77.9012069702148,-77.7494049072266,-77.6988754272461,-77.650993347168,-77.6359710693359,-77.560417175293,-77.559684753418,-77.580078125,-77.5719528198242,-77.5574340820312,-77.5380783081055,-77.4541168212891,-77.4188537597656,-77.4134674072266,-77.3786239624023,-77.389030456543,-77.3349914550781,-77.3094787597656,-77.2935104370117,-77.2704086303711,-77.2559127807617,-77.2574996948242,-77.2410354614258,-77.3098754882812,-77.3322067260742],"lat":[36.0679817199707,35.9947166442871,35.9960632324219,36.0140342712402,36.0263824462891,36.0353851318359,36.0555725097656,36.0823631286621,36.088493347168,36.0973930358887,36.1267013549805,36.1466026306152,36.1441688537598,36.1341133117676,36.1464233398438,36.1442565917969,36.1358184814453,36.1444396972656,36.1567802429199,36.1777305603027,36.1875495910645,36.1837501525879,36.1890411376953,36.1774444580078,36.1959457397461,36.2812309265137,36.3529853820801,36.5006370544434,36.5098876953125,36.4735717773438,36.4789581298828,36.4725875854492,36.4405364990234,36.4063568115234,36.3759460449219,36.3282699584961,36.3144950866699,36.3042144775391,36.3024597167969,36.3197555541992,36.281795501709,36.2558174133301,36.2400856018066,36.2031021118164,36.1828498840332,36.1627655029297,36.1628608703613,36.1555252075195,36.130687713623,36.1184730529785,36.1013031005859,36.0874443054199,36.0679817199707]}]],[[{"lng":[-76.2989273071289,-76.3242340087891,-76.3724212646484,-76.4603500366211,-76.5024642944336,-76.4983444213867,-76.4059677124023,-76.3213653564453,-76.3033599853516,-76.2741317749023,-76.2612838745117,-76.2385330200195,-76.216194152832,-76.1934814453125,-76.1831665039062,-76.2189025878906,-76.1127090454102,-76.1419372558594,-76.234977722168,-76.2989273071289],"lat":[36.2142295837402,36.2336235046387,36.2523460388184,36.3738975524902,36.4522857666016,36.5039024353027,36.4471588134766,36.4096450805664,36.391845703125,36.3814086914062,36.3637580871582,36.3612289428711,36.3278465270996,36.3248519897461,36.3152389526367,36.2966079711914,36.1744194030762,36.1476898193359,36.1633605957031,36.2142295837402]}]],[[{"lng":[-81.0205688476562,-81.0840835571289,-81.1240615844727,-81.1574630737305,-81.2360076904297,-81.3218688964844,-81.3473510742188,-81.3887100219727,-81.3981475830078,-81.4296340942383,-81.4544372558594,-81.5173110961914,-81.5461044311523,-81.4993209838867,-81.4508056640625,-81.4727554321289,-81.4528884887695,-81.431037902832,-81.4123306274414,-81.4063873291016,-81.3674545288086,-81.354133605957,-81.3656921386719,-81.3613739013672,-81.3289947509766,-81.2642440795898,-81.2398910522461,-81.1766738891602,-81.1533660888672,-81.1383972167969,-81.1293792724609,-81.1133117675781,-81.0985641479492,-81.0842514038086,-81.0428009033203,-81.0246429443359,-81.0027770996094,-80.9828414916992,-80.9779510498047,-80.9563903808594,-80.9245681762695,-80.8889236450195,-80.8708648681641,-80.874382019043,-80.8774108886719,-80.9889526367188,-81.0205688476562],"lat":[36.0349349975586,36.0207672119141,36.0312843322754,36.0209808349609,36.0238227844238,35.9893264770508,36.0153579711914,36.0375671386719,36.0560569763184,36.0656623840332,36.0830574035645,36.0954322814941,36.1113929748535,36.1360359191895,36.1906204223633,36.2343559265137,36.2395858764648,36.2607192993164,36.2672920227051,36.2850532531738,36.2786979675293,36.2997169494629,36.3390502929688,36.3531608581543,36.3635025024414,36.3524131774902,36.365364074707,36.4154434204102,36.4247398376465,36.4176254272461,36.4263305664062,36.4228515625,36.43115234375,36.4299201965332,36.4103355407715,36.3778343200684,36.3666801452637,36.3718338012695,36.3913764953613,36.4037971496582,36.3727531433105,36.3544273376465,36.3246231079102,36.2338829040527,36.0524063110352,36.0533485412598,36.0349349975586]}]],[[{"lng":[-81.8062210083008,-81.8171539306641,-81.8223114013672,-81.8502426147461,-81.8851928710938,-81.898567199707,-81.9065628051758,-81.8939971923828,-81.9111557006836,-81.8305740356445,-81.7304916381836,-81.7094573974609,-81.7403793334961,-81.7410736083984,-81.6330642700195,-81.5619812011719,-81.5408401489258,-81.4727554321289,-81.4508056640625,-81.4993209838867,-81.5461044311523,-81.6590042114258,-81.8062210083008],"lat":[36.1045608520508,36.1093864440918,36.1578559875488,36.1814727783203,36.1886367797852,36.1988563537598,36.221866607666,36.2669830322266,36.2907524108887,36.3346557617188,36.3293418884277,36.3337249755859,36.3618583679199,36.3917846679688,36.3406867980957,36.2735939025879,36.2725067138672,36.2343559265137,36.1906204223633,36.1360359191895,36.1113929748535,36.1175918579102,36.1045608520508]}]],[[{"lng":[-76.4805297851562,-76.5369567871094,-76.5755996704102,-76.5944900512695,-76.5710830688477,-76.5625076293945,-76.4603500366211,-76.3724212646484,-76.3242340087891,-76.2989273071289,-76.275505065918,-76.4805297851562],"lat":[36.0797920227051,36.0879249572754,36.1026573181152,36.239818572998,36.2777290344238,36.3405685424805,36.3738975524902,36.2523460388184,36.2336235046387,36.2142295837402,36.1103706359863,36.0797920227051]}]],[[{"lng":[-76.6887359619141,-76.6482162475586,-76.6042404174805,-76.5625076293945,-76.5710830688477,-76.5944900512695,-76.5755996704102,-76.5369567871094,-76.4805297851562,-76.4204254150391,-76.5230102539062,-76.5940017700195,-76.6490173339844,-76.6332092285156,-76.6901550292969,-76.7265090942383,-76.6887359619141],"lat":[36.2945175170898,36.315315246582,36.3149833679199,36.3405685424805,36.2777290344238,36.239818572998,36.1026573181152,36.0879249572754,36.0797920227051,36.0586051940918,36.0071678161621,36.0101318359375,36.0657081604004,36.0371170043945,36.0496101379395,36.1568222045898,36.2945175170898]}]],[[{"lng":[-81.9413452148438,-81.9613952636719,-81.944953918457,-81.947021484375,-81.9883651733398,-82.0050659179688,-82.0491027832031,-82.042724609375,-82.0623245239258,-82.0777587890625,-82.0204544067383,-81.9331130981445,-81.9111557006836,-81.8939971923828,-81.9065628051758,-81.898567199707,-81.8851928710938,-81.8502426147461,-81.8223114013672,-81.8171539306641,-81.8062210083008,-81.7322692871094,-81.8027877807617,-81.8598709106445,-81.8808135986328,-81.9008483886719,-81.9221496582031,-81.9413452148438],"lat":[35.9549751281738,35.9392204284668,35.9186134338379,35.9104118347168,35.9056434631348,35.9144401550293,35.9672355651855,36.0050086975098,36.0355262756348,36.1001396179199,36.129711151123,36.2633209228516,36.2907524108887,36.2669830322266,36.221866607666,36.1988563537598,36.1886367797852,36.1814727783203,36.1578559875488,36.1093864440918,36.1045608520508,36.0584754943848,35.9603309631348,35.9703407287598,35.9895248413086,35.9946517944336,35.9825134277344,35.9549751281738]}]],[[{"lng":[-80.4955444335938,-80.6895751953125,-80.8774108886719,-80.874382019043,-80.7826995849609,-80.7662963867188,-80.7525634765625,-80.7343673706055,-80.7240371704102,-80.6966552734375,-80.6674423217773,-80.6243133544922,-80.5905990600586,-80.5541534423828,-80.5450134277344,-80.5368804931641,-80.4763870239258,-80.4563446044922,-80.4408111572266,-80.4448089599609,-80.4919815063477,-80.5049896240234,-80.5082473754883,-80.4955444335938],"lat":[36.0432662963867,36.0454788208008,36.0524063110352,36.2338829040527,36.2485771179199,36.2618370056152,36.2582969665527,36.2647590637207,36.258472442627,36.259090423584,36.2461013793945,36.2730979919434,36.2682762145996,36.2784309387207,36.2766571044922,36.2567367553711,36.2547264099121,36.2425575256348,36.2194862365723,36.1233062744141,36.1076965332031,36.0949401855469,36.0708847045898,36.0432662963867]}]],[[{"lng":[-78.2545471191406,-78.266845703125,-78.3084106445312,-78.3295440673828,-78.3601150512695,-78.3944778442383,-78.4311447143555,-78.5480270385742,-78.5458526611328,-78.4925231933594,-78.4169540405273,-78.3808517456055,-78.3460464477539,-78.3087615966797,-78.1347198486328,-78.1096267700195,-78.058349609375,-78.0062866210938,-78.1309127807617,-78.2545471191406],"lat":[35.8155250549316,35.8483772277832,35.8934478759766,35.8878479003906,35.9186744689941,35.932300567627,35.9727172851562,36.0141258239746,36.0680885314941,36.1735877990723,36.1621742248535,36.2247505187988,36.2251815795898,36.2600402832031,36.2365837097168,36.213508605957,36.2113227844238,36.1959457397461,36.0214691162109,35.8155250549316]}]],[[{"lng":[-80.0381011962891,-80.1211471557617,-80.2098770141602,-80.2120666503906,-80.3343048095703,-80.3302841186523,-80.380241394043,-80.4115905761719,-80.4179763793945,-80.449951171875,-80.4648666381836,-80.484130859375,-80.4955444335938,-80.5082473754883,-80.5049896240234,-80.4919815063477,-80.4448089599609,-80.4408111572266,-80.4563446044922,-80.453010559082,-80.0256729125977,-80.0381011962891],"lat":[36.0061836242676,36.0215530395508,36.021427154541,35.9901237487793,35.9925651550293,35.9812355041504,35.9679298400879,35.9846115112305,36.0086326599121,36.0298385620117,36.0501937866211,36.0383224487305,36.0432662963867,36.0708847045898,36.0949401855469,36.1076965332031,36.1233062744141,36.2194862365723,36.2425575256348,36.2570877075195,36.2502326965332,36.0061836242676]}]],[[{"lng":[-79.5378189086914,-80.0426025390625,-80.0381011962891,-80.0256729125977,-79.5305786132812,-79.5378189086914],"lat":[35.8909683227539,35.9168128967285,36.0061836242676,36.2502326965332,36.236156463623,35.8909683227539]}]],[[{"lng":[-79.2461929321289,-79.2379913330078,-79.5409851074219,-79.5378189086914,-79.5305786132812,-79.5305099487305,-79.2584991455078,-79.2597732543945,-79.2708206176758,-79.2461929321289],"lat":[35.8681526184082,35.8372459411621,35.8369903564453,35.8909683227539,36.236156463623,36.2461357116699,36.2356872558594,36.0478935241699,35.9046020507812,35.8681526184082]}]],[[{"lng":[-76.7830657958984,-76.8105316162109,-76.8348388671875,-76.8567276000977,-76.8784866333008,-76.8984832763672,-76.9048080444336,-76.9360961914062,-76.9907302856445,-77.0044250488281,-77.0483169555664,-77.0341796875,-77.040885925293,-77.0662994384766,-77.0900115966797,-77.1886749267578,-77.1958923339844,-77.1933975219727,-77.2155380249023,-77.2138061523438,-77.2767181396484,-77.3152542114258,-77.3322067260742,-77.3098754882812,-77.2410354614258,-77.2574996948242,-77.2559127807617,-77.2704086303711,-77.2935104370117,-77.2986145019531,-77.2346115112305,-77.2176666259766,-77.1300735473633,-76.9947509765625,-76.9806900024414,-76.7450637817383,-76.7606735229492,-76.6937637329102,-76.7411270141602,-76.6972198486328,-76.7083206176758,-76.7285995483398,-76.7417221069336,-76.7612609863281,-76.7830657958984],"lat":[35.8592338562012,35.8813095092773,35.8842277526855,35.8357810974121,35.8172912597656,35.8141822814941,35.8679122924805,35.8945999145508,35.8778381347656,35.8618392944336,35.8574905395508,35.9143333435059,35.9317512512207,35.9360084533691,35.9252433776855,35.9312477111816,35.9359550476074,35.988094329834,35.9890670776367,36.0053634643555,36.0276985168457,36.0480842590332,36.0679817199707,36.0874443054199,36.1013031005859,36.1184730529785,36.130687713623,36.1555252075195,36.1628608703613,36.2115287780762,36.214599609375,36.2409820556641,36.2334632873535,36.2355804443359,36.2302360534668,36.2339172363281,36.1445922851562,35.9929695129395,35.9366264343262,35.9415435791016,35.9198341369629,35.9108581542969,35.8830757141113,35.8645439147949,35.8592338562012]}]],[[{"lng":[-79.0181350708008,-79.0953674316406,-79.2461929321289,-79.2708206176758,-79.2597732543945,-79.2584991455078,-79.1592712402344,-78.9510803222656,-79.0181350708008],"lat":[35.8578643798828,35.8539428710938,35.8681526184082,35.9046020507812,36.0478935241699,36.2356872558594,36.2336692810059,36.2338371276855,35.8578643798828]}]],[[{"lng":[-79.0181350708008,-78.9510803222656,-78.8068008422852,-78.8103561401367,-78.8040542602539,-78.7884140014648,-78.7491226196289,-78.7533950805664,-78.7281036376953,-78.7161712646484,-78.7041549682617,-78.7073440551758,-78.7341995239258,-78.7635040283203,-78.8082962036133,-78.8279571533203,-78.9057159423828,-79.0181350708008],"lat":[35.8578643798828,36.2338371276855,36.231575012207,36.114574432373,36.080940246582,36.062183380127,36.063591003418,36.0114631652832,36.0270729064941,36.0264930725098,35.9973258972168,35.9769439697266,35.9345855712891,35.9144744873047,35.9199256896973,35.8602294921875,35.8605155944824,35.8578643798828]}]],[[{"lng":[-78.1869277954102,-78.2056198120117,-78.2115478515625,-78.2340469360352,-78.2545471191406,-78.1309127807617,-78.0062866210938,-77.9851150512695,-77.9733963012695,-77.9558639526367,-77.9392318725586,-77.9224014282227,-77.917839050293,-77.8868408203125,-77.8113021850586,-77.8013687133789,-77.7471694946289,-77.7201843261719,-77.7005081176758,-77.8301467895508,-77.8449249267578,-77.873046875,-78.1869277954102],"lat":[35.7251129150391,35.7253952026367,35.7381935119629,35.745792388916,35.8155250549316,36.0214691162109,36.1959457397461,36.1774444580078,36.1890411376953,36.1837501525879,36.1875495910645,36.1777305603027,36.1567802429199,36.1444396972656,36.1358184814453,36.1442565917969,36.1464233398438,36.1341133117676,36.1441688537598,35.8544960021973,35.8355751037598,35.8447074890137,35.7251129150391]}]],[[{"lng":[-82.1188507080078,-82.1466522216797,-82.1410140991211,-82.1495666503906,-82.1846084594727,-82.1957702636719,-82.1990509033203,-82.2321014404297,-82.2406158447266,-82.2588577270508,-82.3016967773438,-82.3096923828125,-82.3296813964844,-82.3402633666992,-82.3444976806641,-82.3851776123047,-82.4084243774414,-82.3738555908203,-82.3119277954102,-82.2623062133789,-82.2077331542969,-82.154052734375,-82.1180801391602,-82.0777587890625,-82.0623245239258,-82.042724609375,-82.0491027832031,-82.0050659179688,-81.9883651733398,-81.9833450317383,-81.9908981323242,-82.0346832275391,-82.0979919433594,-82.1188507080078],"lat":[35.818531036377,35.8184814453125,35.9002723693848,35.9214477539062,35.9348754882812,35.9473762512207,36.0017700195312,36.0038757324219,35.984203338623,35.9856758117676,36.0293273925781,36.011474609375,36.0142631530762,36.0253982543945,36.070240020752,36.0785064697266,36.0753173828125,36.0986976623535,36.1221504211426,36.1203765869141,36.1470146179199,36.1396217346191,36.0962562561035,36.1001396179199,36.0355262756348,36.0050086975098,35.9672355651855,35.9144401550293,35.9056434631348,35.8875770568848,35.8724746704102,35.8535804748535,35.8438568115234,35.818531036377]}]],[[{"lng":[-77.6712188720703,-77.7331466674805,-77.7574920654297,-77.7550048828125,-77.7671356201172,-77.8301467895508,-77.7005081176758,-77.6856231689453,-77.6423568725586,-77.6062545776367,-77.5478820800781,-77.5366897583008,-77.5358581542969,-77.5251312255859,-77.4639739990234,-77.4382171630859,-77.4257431030273,-77.4053115844727,-77.3423690795898,-77.3581008911133,-77.3968048095703,-77.4056930541992,-77.4185791015625,-77.4269943237305,-77.4547119140625,-77.4732360839844,-77.5055160522461,-77.6712188720703],"lat":[35.6702651977539,35.7395477294922,35.7981033325195,35.824836730957,35.8368644714355,35.8544960021973,36.1441688537598,36.1466026306152,36.1267013549805,36.0973930358887,36.088493347168,36.0823631286621,36.0555725097656,36.0353851318359,36.0263824462891,36.0140342712402,35.9960632324219,35.9947166442871,35.9080047607422,35.8153114318848,35.8284187316895,35.8172645568848,35.8220863342285,35.8081970214844,35.793815612793,35.7996597290039,35.7667541503906,35.6702651977539]}]],[[{"lng":[-81.3281326293945,-81.3766555786133,-81.4175720214844,-81.457145690918,-81.5602569580078,-81.5933685302734,-81.7018280029297,-81.7484436035156,-81.7734756469727,-81.7756881713867,-81.8027877807617,-81.7322692871094,-81.8062210083008,-81.6590042114258,-81.5461044311523,-81.5173110961914,-81.4544372558594,-81.4296340942383,-81.3981475830078,-81.3887100219727,-81.3473510742188,-81.3218688964844,-81.3292617797852,-81.3395690917969,-81.330680847168,-81.3372116088867,-81.3281326293945],"lat":[35.7950592041016,35.7500343322754,35.7559051513672,35.7799339294434,35.775447845459,35.8131256103516,35.8687973022461,35.9207572937012,35.9222030639648,35.9439468383789,35.9603309631348,36.0584754943848,36.1045608520508,36.1175918579102,36.1113929748535,36.0954322814941,36.0830574035645,36.0656623840332,36.0560569763184,36.0375671386719,36.0153579711914,35.9893264770508,35.9892463684082,35.9292411804199,35.8758010864258,35.8276290893555,35.7950592041016]}]],[[{"lng":[-82.2792053222656,-82.2815780639648,-82.3218460083008,-82.3418502807617,-82.3453979492188,-82.3752517700195,-82.4058151245117,-82.4414825439453,-82.4870910644531,-82.4839477539062,-82.5069351196289,-82.4751968383789,-82.4084243774414,-82.3851776123047,-82.3444976806641,-82.3402633666992,-82.3296813964844,-82.3096923828125,-82.3016967773438,-82.2588577270508,-82.2406158447266,-82.2321014404297,-82.1990509033203,-82.1957702636719,-82.1846084594727,-82.1495666503906,-82.1410140991211,-82.1466522216797,-82.1188507080078,-82.1545944213867,-82.1642913818359,-82.211784362793,-82.2792053222656],"lat":[35.6975555419922,35.7202033996582,35.7398490905762,35.7635078430176,35.8051948547363,35.8164024353027,35.8139724731445,35.8822288513184,35.9048843383789,35.9476089477539,35.972541809082,35.9931755065918,36.0753173828125,36.0785064697266,36.070240020752,36.0253982543945,36.0142631530762,36.011474609375,36.0293273925781,35.9856758117676,35.984203338623,36.0038757324219,36.0017700195312,35.9473762512207,35.9348754882812,35.9214477539062,35.9002723693848,35.8184814453125,35.818531036377,35.7983703613281,35.7605247497559,35.7169876098633,35.6975555419922]}]],[[{"lng":[-77.1784591674805,-77.2076644897461,-77.2608184814453,-77.2645034790039,-77.3581008911133,-77.3423690795898,-77.4053115844727,-77.3322067260742,-77.3152542114258,-77.2767181396484,-77.2138061523438,-77.2155380249023,-77.1933975219727,-77.1958923339844,-77.1886749267578,-77.0900115966797,-77.0662994384766,-77.040885925293,-77.0341796875,-77.0483169555664,-77.0044250488281,-76.9907302856445,-76.9360961914062,-76.9048080444336,-76.8984832763672,-76.8784866333008,-76.8567276000977,-76.8348388671875,-76.8105316162109,-76.7830657958984,-76.8066787719727,-76.8052749633789,-76.8259582519531,-76.8383102416992,-76.9799118041992,-77.1612854003906,-77.1784591674805],"lat":[35.7321929931641,35.755126953125,35.759090423584,35.7827796936035,35.8153114318848,35.9080047607422,35.9947166442871,36.0679817199707,36.0480842590332,36.0276985168457,36.0053634643555,35.9890670776367,35.988094329834,35.9359550476074,35.9312477111816,35.9252433776855,35.9360084533691,35.9317512512207,35.9143333435059,35.8574905395508,35.8618392944336,35.8778381347656,35.8945999145508,35.8679122924805,35.8141822814941,35.8172912597656,35.8357810974121,35.8842277526855,35.8813095092773,35.8592338562012,35.8108406066895,35.7903747558594,35.7568817138672,35.7054557800293,35.6501274108887,35.7367782592773,35.7321929931641]}]],[[{"lng":[-78.9210739135742,-78.9988098144531,-78.9388885498047,-78.9444351196289,-78.9057159423828,-78.8279571533203,-78.8082962036133,-78.7635040283203,-78.7341995239258,-78.7073440551758,-78.7041549682617,-78.7161712646484,-78.7281036376953,-78.7533950805664,-78.7491226196289,-78.6955718994141,-78.5480270385742,-78.4311447143555,-78.3944778442383,-78.3601150512695,-78.3295440673828,-78.3084106445312,-78.266845703125,-78.2545471191406,-78.4776382446289,-78.7032165527344,-78.9210739135742],"lat":[35.578857421875,35.6013221740723,35.7614440917969,35.7701148986816,35.8605155944824,35.8602294921875,35.9199256896973,35.9144744873047,35.9345855712891,35.9769439697266,35.9973258972168,36.0264930725098,36.0270729064941,36.0114631652832,36.063591003418,36.0666465759277,36.0141258239746,35.9727172851562,35.932300567627,35.9186744689941,35.8878479003906,35.8934478759766,35.8483772277832,35.8155250549316,35.695613861084,35.5199394226074,35.578857421875]}]],[[{"lng":[-82.8959732055664,-82.8562698364258,-82.8086700439453,-82.7764434814453,-82.7735977172852,-82.7632293701172,-82.6438903808594,-82.628044128418,-82.6044006347656,-82.5922317504883,-82.6058044433594,-82.5993041992188,-82.5541458129883,-82.5069351196289,-82.4839477539062,-82.4870910644531,-82.4414825439453,-82.4058151245117,-82.5005493164062,-82.7663040161133,-82.8056259155273,-82.8432693481445,-82.8811111450195,-82.9075393676758,-82.9521865844727,-82.9430465698242,-82.9627532958984,-82.9068222045898,-82.9140701293945,-82.8959732055664],"lat":[35.9483604431152,35.9474258422852,35.9208717346191,35.9565734863281,35.9875030517578,35.9995460510254,36.0517234802246,36.0543403625488,36.0429878234863,36.0224494934082,36.003547668457,35.9632987976074,35.9561080932617,35.972541809082,35.9476089477539,35.9048843383789,35.8822288513184,35.8139724731445,35.7961273193359,35.6940002441406,35.6849060058594,35.6917266845703,35.6735610961914,35.7278480529785,35.7389984130859,35.7664642333984,35.7918510437012,35.8722152709961,35.9278678894043,35.9483604431152]}]],[[{"lng":[-80.7265167236328,-80.7762298583984,-80.9553451538086,-80.9510726928711,-80.9614334106445,-80.9312744140625,-81.0035781860352,-81.0547790527344,-81.0722045898438,-81.10888671875,-81.0491027832031,-80.9953460693359,-81.0205688476562,-80.9889526367188,-80.8774108886719,-80.6895751953125,-80.7059707641602,-80.7661209106445,-80.7265167236328],"lat":[35.507568359375,35.5068092346191,35.5091209411621,35.5286674499512,35.5435638427734,35.6195907592773,35.6970558166504,35.7134017944336,35.7436485290527,35.771900177002,35.8359680175781,35.9770774841309,36.0349349975586,36.0533485412598,36.0524063110352,36.0454788208008,35.8516578674316,35.6820373535156,35.507568359375]}]],[[{"lng":[-80.4567718505859,-80.4886932373047,-80.526725769043,-80.5723266601562,-80.6075592041016,-80.6349411010742,-80.7059707641602,-80.6895751953125,-80.4955444335938,-80.484130859375,-80.4648666381836,-80.449951171875,-80.4179763793945,-80.4115905761719,-80.380241394043,-80.3630676269531,-80.3617095947266,-80.4059371948242,-80.3865051269531,-80.3864364624023,-80.3954849243164,-80.4148025512695,-80.4260711669922,-80.4465026855469,-80.4577941894531,-80.4794235229492,-80.4728851318359,-80.4506454467773,-80.4567718505859],"lat":[35.7457962036133,35.7756156921387,35.7818069458008,35.8138122558594,35.8222579956055,35.840259552002,35.8516578674316,36.0454788208008,36.0432662963867,36.0383224487305,36.0501937866211,36.0298385620117,36.0086326599121,35.9846115112305,35.9679298400879,35.9421234130859,35.8958511352539,35.8843688964844,35.858570098877,35.8454132080078,35.839485168457,35.8448638916016,35.8303070068359,35.8306846618652,35.8224754333496,35.8332786560059,35.7861175537109,35.7648773193359,35.7457962036133]}]],[[{"lng":[-81.10888671875,-81.1272811889648,-81.1414031982422,-81.3281326293945,-81.3372116088867,-81.330680847168,-81.3395690917969,-81.3292617797852,-81.3218688964844,-81.2360076904297,-81.1574630737305,-81.1240615844727,-81.0840835571289,-81.0205688476562,-80.9953460693359,-81.0491027832031,-81.10888671875],"lat":[35.771900177002,35.7889671325684,35.8233184814453,35.7950592041016,35.8276290893555,35.8758010864258,35.9292411804199,35.9892463684082,35.9893264770508,36.0238227844238,36.0209808349609,36.0312843322754,36.0207672119141,36.0349349975586,35.9770774841309,35.8359680175781,35.771900177002]}]],[[{"lng":[-80.0644073486328,-80.1819076538086,-80.2092132568359,-80.251220703125,-80.2857818603516,-80.3255920410156,-80.326286315918,-80.3772583007812,-80.4567718505859,-80.4506454467773,-80.4728851318359,-80.4794235229492,-80.4577941894531,-80.4465026855469,-80.4260711669922,-80.4148025512695,-80.3954849243164,-80.3864364624023,-80.3865051269531,-80.4059371948242,-80.3617095947266,-80.3630676269531,-80.380241394043,-80.3302841186523,-80.3343048095703,-80.2120666503906,-80.2098770141602,-80.1211471557617,-80.0381011962891,-80.0426025390625,-80.0644073486328],"lat":[35.5056991577148,35.5051193237305,35.5745124816895,35.6248016357422,35.6369743347168,35.6831550598145,35.7140121459961,35.7106857299805,35.7457962036133,35.7648773193359,35.7861175537109,35.8332786560059,35.8224754333496,35.8306846618652,35.8303070068359,35.8448638916016,35.839485168457,35.8454132080078,35.858570098877,35.8843688964844,35.8958511352539,35.9421234130859,35.9679298400879,35.9812355041504,35.9925651550293,35.9901237487793,36.021427154541,36.0215530395508,36.0061836242676,35.9168128967285,35.5056991577148]}]],[[{"lng":[-81.8162841796875,-81.8656234741211,-81.985595703125,-81.9752960205078,-81.9329833984375,-81.9062576293945,-81.9062271118164,-81.9413452148438,-81.9221496582031,-81.9008483886719,-81.8808135986328,-81.8598709106445,-81.8027877807617,-81.7756881713867,-81.7734756469727,-81.7484436035156,-81.7018280029297,-81.5933685302734,-81.5602569580078,-81.457145690918,-81.4175720214844,-81.3766555786133,-81.4057235717773,-81.4956970214844,-81.5236434936523,-81.5625381469727,-81.5881118774414,-81.6439056396484,-81.6843185424805,-81.695198059082,-81.7008209228516,-81.7496109008789,-81.7656097412109,-81.8162841796875],"lat":[35.5743789672852,35.7188529968262,35.7999534606934,35.8182792663574,35.8275985717773,35.8479957580566,35.8920135498047,35.9549751281738,35.9825134277344,35.9946517944336,35.9895248413086,35.9703407287598,35.9603309631348,35.9439468383789,35.9222030639648,35.9207572937012,35.8687973022461,35.8131256103516,35.775447845459,35.7799339294434,35.7559051513672,35.7500343322754,35.697509765625,35.6065483093262,35.5612602233887,35.5553092956543,35.5617752075195,35.5532989501953,35.5654373168945,35.5716361999512,35.5956115722656,35.6017112731934,35.5842247009277,35.5743789672852]}]],[[{"lng":[-76.4084320068359,-76.6338195800781,-76.8383102416992,-76.8259582519531,-76.8052749633789,-76.8066787719727,-76.7830657958984,-76.7612609863281,-76.7417221069336,-76.7285995483398,-76.7083206176758,-76.6972198486328,-76.4094696044922,-76.3714828491211,-76.3835601806641,-76.3581924438477,-76.4086990356445,-76.4084320068359],"lat":[35.6991157531738,35.7030029296875,35.7054557800293,35.7568817138672,35.7903747558594,35.8108406066895,35.8592338562012,35.8645439147949,35.8830757141113,35.9108581542969,35.9198341369629,35.9415435791016,35.977466583252,35.9323425292969,35.9004135131836,35.8605918884277,35.7894821166992,35.6991157531738]}]],[[{"lng":[-76.1673049926758,-76.2102355957031,-76.232795715332,-76.2976303100586,-76.2734451293945,-76.4084320068359,-76.4086990356445,-76.3581924438477,-76.3835601806641,-76.3714828491211,-76.2137680053711,-76.0896377563477,-76.0260467529297,-76.0759124755859,-76.0430679321289,-76.1673049926758],"lat":[35.6968421936035,35.6043891906738,35.5946655273438,35.6116943359375,35.6894989013672,35.6991157531738,35.7894821166992,35.8605918884277,35.9004135131836,35.9323425292969,35.9768753051758,35.9629135131836,35.9204254150391,35.7568016052246,35.6838493347168,35.6968421936035]}]],[[{"lng":[-81.8162841796875,-81.8318939208984,-81.8402709960938,-81.8621520996094,-81.9670486450195,-82.0016021728516,-82.0361862182617,-82.1167526245117,-82.1707077026367,-82.2680053710938,-82.2906723022461,-82.2653427124023,-82.2842407226562,-82.2891845703125,-82.2792053222656,-82.211784362793,-82.1642913818359,-82.1545944213867,-82.1188507080078,-82.0979919433594,-82.0346832275391,-81.9908981323242,-81.9833450317383,-81.9883651733398,-81.947021484375,-81.944953918457,-81.9613952636719,-81.9413452148438,-81.9062271118164,-81.9062576293945,-81.9329833984375,-81.9752960205078,-81.985595703125,-81.8656234741211,-81.8162841796875],"lat":[35.5743789672852,35.5650634765625,35.5372428894043,35.5305480957031,35.5215873718262,35.5459747314453,35.5285873413086,35.5190048217773,35.5284652709961,35.5688591003418,35.5893020629883,35.6124877929688,35.6389045715332,35.6714897155762,35.6975555419922,35.7169876098633,35.7605247497559,35.7983703613281,35.818531036377,35.8438568115234,35.8535804748535,35.8724746704102,35.8875770568848,35.9056434631348,35.9104118347168,35.9186134338379,35.9392204284668,35.9549751281738,35.8920135498047,35.8479957580566,35.8275985717773,35.8182792663574,35.7999534606934,35.7188529968262,35.5743789672852]}]],[[{"lng":[-79.7649917602539,-80.0644073486328,-80.0426025390625,-79.5378189086914,-79.5409851074219,-79.5553588867188,-79.7649917602539],"lat":[35.5059356689453,35.5056991577148,35.9168128967285,35.8909683227539,35.8369903564453,35.5130500793457,35.5059356689453]}]],[[{"lng":[-79.5553588867188,-79.5409851074219,-79.2379913330078,-79.2461929321289,-79.0953674316406,-79.0181350708008,-78.9057159423828,-78.9444351196289,-78.9388885498047,-78.9988098144531,-78.9210739135742,-78.9744873046875,-79.0384979248047,-79.0672912597656,-79.1326065063477,-79.1942443847656,-79.2097320556641,-79.2205047607422,-79.2271347045898,-79.2760925292969,-79.2861328125,-79.3053665161133,-79.3150100708008,-79.312858581543,-79.3315505981445,-79.3429489135742,-79.5553588867188],"lat":[35.5130500793457,35.8369903564453,35.8372459411621,35.8681526184082,35.8539428710938,35.8578643798828,35.8605155944824,35.7701148986816,35.7614440917969,35.6013221740723,35.578857421875,35.5171546936035,35.5494613647461,35.598705291748,35.6246147155762,35.5751304626465,35.5534515380859,35.5508003234863,35.5653648376465,35.5302772521973,35.5444068908691,35.5427055358887,35.5391311645508,35.5268669128418,35.5219802856445,35.5102424621582,35.5130500793457]}]],[[{"lng":[-78.0653305053711,-78.1249008178711,-78.1624450683594,-78.1618041992188,-78.1869277954102,-77.873046875,-77.8449249267578,-77.8301467895508,-77.7671356201172,-77.7550048828125,-77.7574920654297,-77.7331466674805,-77.6712188720703,-77.6983261108398,-77.8262939453125,-77.8271789550781,-78.0021362304688,-78.0604858398438,-78.0653305053711],"lat":[35.5820388793945,35.5975189208984,35.6829833984375,35.7092933654785,35.7251129150391,35.8447074890137,35.8355751037598,35.8544960021973,35.8368644714355,35.824836730957,35.7981033325195,35.7395477294922,35.6702651977539,35.6539993286133,35.5742568969727,35.5829010009766,35.5759963989258,35.594669342041,35.5820388793945]}]],[[{"lng":[-80.2982406616211,-80.7265167236328,-80.7661209106445,-80.7059707641602,-80.6349411010742,-80.6075592041016,-80.5723266601562,-80.526725769043,-80.4886932373047,-80.4567718505859,-80.3772583007812,-80.326286315918,-80.3255920410156,-80.2857818603516,-80.251220703125,-80.2092132568359,-80.1819076538086,-80.2982406616211],"lat":[35.4948997497559,35.507568359375,35.6820373535156,35.8516578674316,35.840259552002,35.8222579956055,35.8138122558594,35.7818069458008,35.7756156921387,35.7457962036133,35.7106857299805,35.7140121459961,35.6831550598145,35.6369743347168,35.6248016357422,35.5745124816895,35.5051193237305,35.4948997497559]}]],[[{"lng":[-77.4738845825195,-77.5045623779297,-77.5039367675781,-77.5210494995117,-77.5234298706055,-77.5495758056641,-77.5646438598633,-77.6341323852539,-77.6983261108398,-77.6712188720703,-77.5055160522461,-77.4732360839844,-77.4547119140625,-77.4269943237305,-77.4185791015625,-77.4056930541992,-77.3968048095703,-77.3581008911133,-77.2645034790039,-77.2608184814453,-77.2076644897461,-77.1784591674805,-77.1774215698242,-77.1922378540039,-77.1955184936523,-77.1754379272461,-77.1872100830078,-77.1746826171875,-77.1520538330078,-77.1483459472656,-77.1193923950195,-77.1037673950195,-77.1473999023438,-77.1722106933594,-77.1949615478516,-77.2112045288086,-77.2405471801758,-77.2443008422852,-77.2641906738281,-77.2938003540039,-77.3542175292969,-77.3861618041992,-77.4014892578125,-77.4439468383789,-77.4738845825195],"lat":[35.4215278625488,35.4848327636719,35.5038909912109,35.5165061950684,35.5301780700684,35.5257263183594,35.5319404602051,35.5878257751465,35.6539993286133,35.6702651977539,35.7667541503906,35.7996597290039,35.793815612793,35.8081970214844,35.8220863342285,35.8172645568848,35.8284187316895,35.8153114318848,35.7827796936035,35.759090423584,35.755126953125,35.7321929931641,35.7144622802734,35.7120933532715,35.6999130249023,35.6762847900391,35.664306640625,35.6354103088379,35.6198844909668,35.5980033874512,35.5854988098145,35.5501861572266,35.5475883483887,35.5191230773926,35.4229545593262,35.3956413269043,35.3799858093262,35.354190826416,35.3501129150391,35.373950958252,35.3235549926758,35.3292617797852,35.3427696228027,35.3545951843262,35.4215278625488]}]],[[{"lng":[-80.9614334106445,-81.5236434936523,-81.4956970214844,-81.4057235717773,-81.3766555786133,-81.3281326293945,-81.1414031982422,-81.1272811889648,-81.10888671875,-81.0722045898438,-81.0547790527344,-81.0035781860352,-80.9312744140625,-80.9614334106445],"lat":[35.5435638427734,35.5612602233887,35.6065483093262,35.697509765625,35.7500343322754,35.7950592041016,35.8233184814453,35.7889671325684,35.771900177002,35.7436485290527,35.7134017944336,35.6970558166504,35.6195907592773,35.5435638427734]}]],[[{"lng":[-82.2581024169922,-82.3228759765625,-82.370964050293,-82.3648376464844,-82.3736877441406,-82.409065246582,-82.4749221801758,-82.5195159912109,-82.5269012451172,-82.5511322021484,-82.5639343261719,-82.6186828613281,-82.6689834594727,-82.7141571044922,-82.7438888549805,-82.7808837890625,-82.7947235107422,-82.7706298828125,-82.7720336914062,-82.8133392333984,-82.8231048583984,-82.8448638916016,-82.8655242919922,-82.8811111450195,-82.8432693481445,-82.8056259155273,-82.7663040161133,-82.5005493164062,-82.4058151245117,-82.3752517700195,-82.3453979492188,-82.3418502807617,-82.3218460083008,-82.2815780639648,-82.2792053222656,-82.2891845703125,-82.2842407226562,-82.2653427124023,-82.2906723022461,-82.2680053710938,-82.1707077026367,-82.2229080200195,-82.2276000976562,-82.2401580810547,-82.2581024169922],"lat":[35.4637298583984,35.4951591491699,35.4787712097168,35.4634590148926,35.4573783874512,35.468921661377,35.4444046020508,35.4439163208008,35.4283218383789,35.4268951416016,35.4384231567383,35.4376754760742,35.4551658630371,35.439151763916,35.4180335998535,35.4416885375977,35.4649696350098,35.5327339172363,35.5712928771973,35.6175346374512,35.6213874816895,35.6131477355957,35.6358032226562,35.6735610961914,35.6917266845703,35.6849060058594,35.6940002441406,35.7961273193359,35.8139724731445,35.8164024353027,35.8051948547363,35.7635078430176,35.7398490905762,35.7202033996582,35.6975555419922,35.6714897155762,35.6389045715332,35.6124877929688,35.5893020629883,35.5688591003418,35.5284652709961,35.5156936645508,35.4829216003418,35.4681549072266,35.4637298583984]}]],[[{"lng":[-78.5387420654297,-78.5394668579102,-78.6008224487305,-78.623176574707,-78.6887435913086,-78.7089691162109,-78.7032165527344,-78.4776382446289,-78.2545471191406,-78.2340469360352,-78.2115478515625,-78.2056198120117,-78.1869277954102,-78.1618041992188,-78.1624450683594,-78.1249008178711,-78.0653305053711,-78.1459732055664,-78.1568908691406,-78.1774597167969,-78.2090377807617,-78.2372283935547,-78.2681655883789,-78.3101272583008,-78.4166793823242,-78.4936218261719,-78.5387420654297],"lat":[35.3151168823242,35.3364639282227,35.4030303955078,35.4464149475098,35.5075263977051,35.514102935791,35.5199394226074,35.695613861084,35.8155250549316,35.745792388916,35.7381935119629,35.7253952026367,35.7251129150391,35.7092933654785,35.6829833984375,35.5975189208984,35.5820388793945,35.4289932250977,35.3506317138672,35.339599609375,35.3391761779785,35.3146324157715,35.3173599243164,35.2802848815918,35.2490768432617,35.2596206665039,35.3151168823242]}]],[[{"lng":[-82.7438888549805,-82.8332366943359,-82.9191131591797,-82.9536209106445,-82.9853591918945,-83.0385971069336,-83.0460433959961,-83.0877151489258,-83.1282348632812,-83.1423721313477,-83.181396484375,-83.1570205688477,-83.1803741455078,-83.1781005859375,-83.1951293945312,-83.1842956542969,-83.2152557373047,-83.2591247558594,-83.253303527832,-83.2438507080078,-83.1853485107422,-83.1436614990234,-83.1181869506836,-83.0599594116211,-82.9870071411133,-82.9627532958984,-82.9430465698242,-82.9521865844727,-82.9075393676758,-82.8811111450195,-82.8655242919922,-82.8448638916016,-82.8231048583984,-82.8133392333984,-82.7720336914062,-82.7706298828125,-82.7947235107422,-82.7808837890625,-82.7438888549805],"lat":[35.4180335998535,35.3155708312988,35.2907676696777,35.3085289001465,35.3563232421875,35.3899307250977,35.4069938659668,35.4463272094727,35.456615447998,35.4843940734863,35.5124092102051,35.5516624450684,35.582820892334,35.623291015625,35.6378173828125,35.6630783081055,35.6853981018066,35.6910095214844,35.7007064819336,35.7182159423828,35.7288856506348,35.7626838684082,35.7638092041016,35.7825775146484,35.7739906311035,35.7918510437012,35.7664642333984,35.7389984130859,35.7278480529785,35.6735610961914,35.6358032226562,35.6131477355957,35.6213874816895,35.6175346374512,35.5712928771973,35.5327339172363,35.4649696350098,35.4416885375977,35.4180335998535]}]],[[{"lng":[-75.7831726074219,-75.7731552124023,-75.5449676513672,-75.7027359008789,-75.7408676147461,-75.7831726074219],"lat":[36.2251930236816,36.2292556762695,35.7883605957031,36.049861907959,36.0503234863281,36.2251930236816]}],[{"lng":[-75.8914947509766,-75.9080276489258,-76.0212097167969,-75.9878540039062,-75.8180541992188,-75.7489624023438,-75.7293701171875,-75.779052734375,-75.8914947509766],"lat":[35.6312675476074,35.6656379699707,35.6690940856934,35.892707824707,35.9235191345215,35.8693389892578,35.6651725769043,35.578685760498,35.6312675476074]}],[{"lng":[-75.4912185668945,-75.5336227416992,-75.4569778442383,-75.5262985229492,-75.7492904663086,-75.6915664672852,-75.521484375,-75.4754180908203,-75.4912185668945],"lat":[35.6704978942871,35.768856048584,35.6173973083496,35.2279167175293,35.189826965332,35.2349891662598,35.2813568115234,35.5644950866699,35.6704978942871]}]],[[{"lng":[-77.1037673950195,-77.1193923950195,-77.1483459472656,-77.1520538330078,-77.1746826171875,-77.1872100830078,-77.1754379272461,-77.1955184936523,-77.1922378540039,-77.1774215698242,-77.1784591674805,-77.1612854003906,-76.9799118041992,-76.8383102416992,-76.6338195800781,-76.6089172363281,-76.6079483032227,-76.5858764648438,-76.5395965576172,-76.5189437866211,-76.4925384521484,-76.6381988525391,-76.6287689208984,-76.7053756713867,-77.1037673950195],"lat":[35.5501861572266,35.5854988098145,35.5980033874512,35.6198844909668,35.6354103088379,35.664306640625,35.6762847900391,35.6999130249023,35.7120933532715,35.7144622802734,35.7321929931641,35.7367782592773,35.6501274108887,35.7054557800293,35.7030029296875,35.6641540527344,35.635066986084,35.6094551086426,35.5940361022949,35.5776443481445,35.5417861938477,35.520336151123,35.4378967285156,35.4119338989258,35.5501861572266]}],[{"lng":[-76.6145172119141,-76.6402130126953,-76.8505706787109,-76.8976058959961,-77.1949615478516,-77.1722106933594,-77.1473999023438,-77.1037673950195,-76.9831848144531,-76.6949005126953,-76.6145172119141],"lat":[35.2729187011719,35.237247467041,35.2171669006348,35.2515716552734,35.4229545593262,35.5191230773926,35.5475883483887,35.5501861572266,35.4365005493164,35.3504257202148,35.2729187011719]}]],[[{"lng":[-83.3318176269531,-83.4246826171875,-83.4732284545898,-83.6791915893555,-83.6900863647461,-83.6893844604492,-83.6342163085938,-83.5988845825195,-83.5803833007812,-83.5867156982422,-83.63232421875,-83.6536102294922,-83.7449569702148,-83.8549880981445,-83.8798904418945,-83.9300765991211,-83.954704284668,-83.9100112915039,-83.8812255859375,-83.8302001953125,-83.77587890625,-83.6728744506836,-83.6138610839844,-83.56103515625,-83.5057983398438,-83.4582901000977,-83.387092590332,-83.3430252075195,-83.2984161376953,-83.2591247558594,-83.2152557373047,-83.1842956542969,-83.1951293945312,-83.1781005859375,-83.1803741455078,-83.1570205688477,-83.181396484375,-83.2266616821289,-83.2473068237305,-83.3098602294922,-83.35302734375,-83.3647155761719,-83.3298110961914,-83.3318176269531],"lat":[35.3193435668945,35.3153190612793,35.2916450500488,35.2722206115723,35.2877807617188,35.3082427978516,35.3340339660645,35.3623657226562,35.4010848999023,35.4276885986328,35.4367408752441,35.4210968017578,35.4414024353027,35.4501266479492,35.462028503418,35.4490165710449,35.4554595947266,35.4764785766602,35.5105857849121,35.5190620422363,35.552604675293,35.5649719238281,35.5717391967773,35.55517578125,35.5595512390137,35.5972785949707,35.6252174377441,35.6532592773438,35.6563262939453,35.6910095214844,35.6853981018066,35.6630783081055,35.6378173828125,35.623291015625,35.582820892334,35.5516624450684,35.5124092102051,35.5134544372559,35.5069885253906,35.4634742736816,35.4558982849121,35.413330078125,35.3639106750488,35.3193435668945]}]],[[{"lng":[-77.8051834106445,-77.8041076660156,-77.8305816650391,-77.8262939453125,-77.6983261108398,-77.6341323852539,-77.5646438598633,-77.5495758056641,-77.5234298706055,-77.5210494995117,-77.5039367675781,-77.5045623779297,-77.4738845825195,-77.4946746826172,-77.5349731445312,-77.5371780395508,-77.558723449707,-77.6218032836914,-77.6660537719727,-77.6844711303711,-77.6984176635742,-77.7606964111328,-77.8051834106445],"lat":[35.3645896911621,35.401798248291,35.4236259460449,35.5742568969727,35.6539993286133,35.5878257751465,35.5319404602051,35.5257263183594,35.5301780700684,35.5165061950684,35.5038909912109,35.4848327636719,35.4215278625488,35.4074440002441,35.4178237915039,35.4019813537598,35.3810882568359,35.3664855957031,35.339672088623,35.3459396362305,35.3698196411133,35.3619384765625,35.3645896911621]}]],[[{"lng":[-79.1824417114258,-79.2643203735352,-79.2847290039062,-79.3031997680664,-79.3245849609375,-79.3310928344727,-79.3624877929688,-79.3436737060547,-79.3429489135742,-79.3315505981445,-79.312858581543,-79.3150100708008,-79.3053665161133,-79.2861328125,-79.2760925292969,-79.2271347045898,-79.2205047607422,-79.2097320556641,-79.1942443847656,-79.1326065063477,-79.0672912597656,-79.0384979248047,-78.9744873046875,-79.1824417114258],"lat":[35.3045387268066,35.3445701599121,35.3928070068359,35.408805847168,35.4148292541504,35.4448204040527,35.4704132080078,35.491641998291,35.5102424621582,35.5219802856445,35.5268669128418,35.5391311645508,35.5427055358887,35.5444068908691,35.5302772521973,35.5653648376465,35.5508003234863,35.5534515380859,35.5751304626465,35.6246147155762,35.598705291748,35.5494613647461,35.5171546936035,35.3045387268066]}]],[[{"lng":[-81.9714431762695,-81.9645156860352,-82.0515823364258,-82.0841522216797,-82.1221694946289,-82.1551284790039,-82.2582778930664,-82.2787246704102,-82.2746810913086,-82.2581024169922,-82.2401580810547,-82.2276000976562,-82.2229080200195,-82.1707077026367,-82.1167526245117,-82.0361862182617,-82.0016021728516,-81.9670486450195,-81.8621520996094,-81.8402709960938,-81.8318939208984,-81.8162841796875,-81.7656097412109,-81.7496109008789,-81.7008209228516,-81.695198059082,-81.6843185424805,-81.6976852416992,-81.7371673583984,-81.7594909667969,-81.7653579711914,-81.8705902099609,-81.9714431762695],"lat":[35.1882820129395,35.2474250793457,35.323127746582,35.33935546875,35.3886184692383,35.397102355957,35.3933563232422,35.4356422424316,35.4516067504883,35.4637298583984,35.4681549072266,35.4829216003418,35.5156936645508,35.5284652709961,35.5190048217773,35.5285873413086,35.5459747314453,35.5215873718262,35.5305480957031,35.5372428894043,35.5650634765625,35.5743789672852,35.5842247009277,35.6017112731934,35.5956115722656,35.5716361999512,35.5654373168945,35.3532638549805,35.2541732788086,35.2206993103027,35.1824722290039,35.1831169128418,35.1882820129395]}]],[[{"lng":[-78.1631927490234,-78.1651763916016,-78.2574005126953,-78.3101272583008,-78.2681655883789,-78.2372283935547,-78.2090377807617,-78.1774597167969,-78.1568908691406,-78.1459732055664,-78.0653305053711,-78.0604858398438,-78.0021362304688,-77.8271789550781,-77.8262939453125,-77.8305816650391,-77.8041076660156,-77.8051834106445,-77.8300628662109,-77.8365783691406,-77.8876724243164,-77.894157409668,-77.9139785766602,-77.944694519043,-77.9639282226562,-78.0021591186523,-78.036506652832,-78.087532043457,-78.1631927490234],"lat":[35.1822891235352,35.1932182312012,35.2209510803223,35.2802848815918,35.3173599243164,35.3146324157715,35.3391761779785,35.339599609375,35.3506317138672,35.4289932250977,35.5820388793945,35.594669342041,35.5759963989258,35.5829010009766,35.5742568969727,35.4236259460449,35.401798248291,35.3645896911621,35.3423500061035,35.1717414855957,35.1549644470215,35.1441802978516,35.1599731445312,35.1682319641113,35.1640243530273,35.1864776611328,35.1856918334961,35.1701850891113,35.1822891235352]}]],[[{"lng":[-78.6127395629883,-78.716064453125,-78.812385559082,-78.8745727539062,-78.8860168457031,-78.9125823974609,-79.0958938598633,-79.1468353271484,-79.1690979003906,-79.2166519165039,-79.1824417114258,-78.9744873046875,-78.9210739135742,-78.7032165527344,-78.7089691162109,-78.6887435913086,-78.623176574707,-78.6008224487305,-78.5394668579102,-78.5387420654297,-78.5804138183594,-78.6127395629883],"lat":[35.2438316345215,35.2599792480469,35.2587203979492,35.2425346374512,35.2299346923828,35.2224655151367,35.1899642944336,35.2130432128906,35.2359008789062,35.2652778625488,35.3045387268066,35.5171546936035,35.578857421875,35.5199394226074,35.514102935791,35.5075263977051,35.4464149475098,35.4030303955078,35.3364639282227,35.3151168823242,35.2865753173828,35.2438316345215]}]],[[{"lng":[-81.3228225708008,-81.362174987793,-81.7653579711914,-81.7594909667969,-81.7371673583984,-81.6976852416992,-81.6843185424805,-81.6439056396484,-81.5881118774414,-81.5625381469727,-81.5236434936523,-81.5069580078125,-81.5155029296875,-81.4450759887695,-81.4074172973633,-81.3532638549805,-81.3647994995117,-81.3523483276367,-81.3194274902344,-81.3114242553711,-81.3228225708008],"lat":[35.1637573242188,35.1628532409668,35.1824722290039,35.2206993103027,35.2541732788086,35.3532638549805,35.5654373168945,35.5532989501953,35.5617752075195,35.5553092956543,35.5612602233887,35.5464973449707,35.5214233398438,35.4133682250977,35.359806060791,35.3273010253906,35.3103713989258,35.2751045227051,35.260498046875,35.1879501342773,35.1637573242188]}]],[[{"lng":[-80.9567718505859,-81.4450759887695,-81.5155029296875,-81.5069580078125,-81.5236434936523,-80.9614334106445,-80.9510726928711,-80.9553451538086,-80.9421997070312,-80.9538650512695,-80.9567718505859],"lat":[35.3974533081055,35.4133682250977,35.5214233398438,35.5464973449707,35.5612602233887,35.5435638427734,35.5286674499512,35.5091209411621,35.4511299133301,35.4360580444336,35.3974533081055]}]],[[{"lng":[-83.1062850952148,-83.1614990234375,-83.1450576782227,-83.1482009887695,-83.1771469116211,-83.1934814453125,-83.2179336547852,-83.2200393676758,-83.232780456543,-83.2639389038086,-83.2849807739258,-83.3106384277344,-83.3318176269531,-83.3298110961914,-83.3647155761719,-83.35302734375,-83.3098602294922,-83.2473068237305,-83.2266616821289,-83.181396484375,-83.1423721313477,-83.1282348632812,-83.0877151489258,-83.0460433959961,-83.0385971069336,-82.9853591918945,-82.9536209106445,-82.9191131591797,-82.9268569946289,-82.9787139892578,-82.9905853271484,-82.9820785522461,-83.0520858764648,-83.0438537597656,-83.0072784423828,-83.1062850952148],"lat":[35.0002784729004,35.0592231750488,35.0832862854004,35.091381072998,35.1114959716797,35.139217376709,35.158992767334,35.222526550293,35.2308082580566,35.2217712402344,35.226634979248,35.2599868774414,35.3193435668945,35.3639106750488,35.413330078125,35.4558982849121,35.4634742736816,35.5069885253906,35.5134544372559,35.5124092102051,35.4843940734863,35.456615447998,35.4463272094727,35.4069938659668,35.3899307250977,35.3563232421875,35.3085289001465,35.2907676696777,35.2279052734375,35.1857299804688,35.1581764221191,35.1275062561035,35.0634765625,35.0400695800781,35.0242042541504,35.0002784729004]}]],[[{"lng":[-79.6074676513672,-79.6483764648438,-79.7082366943359,-79.7649917602539,-79.5553588867188,-79.3429489135742,-79.3436737060547,-79.3624877929688,-79.3310928344727,-79.3245849609375,-79.3031997680664,-79.2847290039062,-79.2643203735352,-79.1824417114258,-79.2166519165039,-79.1690979003906,-79.1468353271484,-79.0958938598633,-79.0971755981445,-79.1220016479492,-79.1472778320312,-79.2352905273438,-79.3500518798828,-79.4553604125977,-79.4950637817383,-79.5529022216797,-79.5741958618164,-79.5756301879883,-79.6074676513672],"lat":[35.1591873168945,35.1897315979004,35.2784156799316,35.5059356689453,35.5130500793457,35.5102424621582,35.491641998291,35.4704132080078,35.4448204040527,35.4148292541504,35.408805847168,35.3928070068359,35.3445701599121,35.3045387268066,35.2652778625488,35.2359008789062,35.2130432128906,35.1899642944336,35.1768074035645,35.1701850891113,35.1735496520996,35.2032089233398,35.140796661377,35.0373573303223,35.0625076293945,35.0641059875488,35.0732650756836,35.1191253662109,35.1591873168945]}]],[[{"lng":[-81.0493011474609,-81.0239639282227,-81.0072784423828,-81.0015182495117,-81.0140533447266,-80.9796447753906,-80.9262771606445,-80.9377593994141,-80.9829254150391,-80.9567718505859,-80.9538650512695,-80.9421997070312,-80.9553451538086,-80.7762298583984,-80.7753829956055,-80.7616882324219,-80.7454376220703,-80.7627410888672,-80.6899871826172,-80.6594467163086,-80.5396423339844,-80.7596817016602,-80.7972183227539,-80.8401641845703,-80.8947143554688,-80.9277954101562,-81.0398864746094,-81.0655517578125,-81.0284423828125,-81.0490417480469,-81.0493011474609],"lat":[35.1515312194824,35.1490325927734,35.1632499694824,35.195987701416,35.2499008178711,35.3332672119141,35.3486747741699,35.3649253845215,35.3690948486328,35.3974533081055,35.4360580444336,35.4511299133301,35.5091209411621,35.5068092346191,35.4795722961426,35.4651374816895,35.4198570251465,35.4006805419922,35.3407592773438,35.2646751403809,35.2058181762695,35.03662109375,35.0281982421875,35.0020179748535,35.0597343444824,35.1012496948242,35.0372009277344,35.0664825439453,35.1054077148438,35.1320114135742,35.1515312194824]}]],[[{"lng":[-80.5029449462891,-80.5396423339844,-80.6594467163086,-80.6899871826172,-80.7627410888672,-80.7454376220703,-80.7616882324219,-80.7753829956055,-80.7762298583984,-80.7265167236328,-80.2982406616211,-80.3784713745117,-80.4817199707031,-80.5029449462891],"lat":[35.1869125366211,35.2058181762695,35.2646751403809,35.3407592773438,35.4006805419922,35.4198570251465,35.4651374816895,35.4795722961426,35.5068092346191,35.507568359375,35.4948997497559,35.384838104248,35.2106094360352,35.1869125366211]}]],[[{"lng":[-80.0714111328125,-80.0590667724609,-80.0658416748047,-80.0923461914062,-80.0935134887695,-80.0507507324219,-80.051887512207,-80.1163101196289,-80.1620712280273,-80.1819076538086,-80.0644073486328,-79.7649917602539,-79.7082366943359,-79.6483764648438,-79.6074676513672,-79.6372985839844,-79.6979522705078,-79.7598419189453,-79.8363189697266,-79.9094619750977,-79.9758224487305,-80.0106811523438,-80.0651779174805,-80.0714111328125],"lat":[35.1507415771484,35.1770782470703,35.2079429626465,35.2542266845703,35.291446685791,35.3663711547852,35.3768081665039,35.4643592834473,35.4747428894043,35.5051193237305,35.5056991577148,35.5059356689453,35.2784156799316,35.1897315979004,35.1591873168945,35.1538352966309,35.1730728149414,35.1673126220703,35.1738014221191,35.1557197570801,35.1507568359375,35.1371421813965,35.1365814208984,35.1507415771484]}]],[[{"lng":[-80.0714111328125,-80.1102752685547,-80.1316146850586,-80.1546859741211,-80.1591415405273,-80.1681289672852,-80.1901016235352,-80.2198867797852,-80.2475891113281,-80.2610931396484,-80.2751235961914,-80.3256759643555,-80.3487243652344,-80.3729476928711,-80.3987503051758,-80.428581237793,-80.4532852172852,-80.4832305908203,-80.5029449462891,-80.4817199707031,-80.3784713745117,-80.2982406616211,-80.1819076538086,-80.1620712280273,-80.1163101196289,-80.051887512207,-80.0507507324219,-80.0935134887695,-80.0923461914062,-80.0658416748047,-80.0590667724609,-80.0714111328125],"lat":[35.1507415771484,35.1938362121582,35.1711158752441,35.176082611084,35.1547393798828,35.149730682373,35.1678581237793,35.1591835021973,35.2045249938965,35.2040405273438,35.1931114196777,35.1739158630371,35.1706733703613,35.177864074707,35.1646118164062,35.1685943603516,35.1607856750488,35.1829109191895,35.1869125366211,35.2106094360352,35.384838104248,35.4948997497559,35.5051193237305,35.4747428894043,35.4643592834473,35.3768081665039,35.3663711547852,35.291446685791,35.2542266845703,35.2079429626465,35.1770782470703,35.1507415771484]}]],[[{"lng":[-82.5700302124023,-82.589225769043,-82.5962371826172,-82.6106109619141,-82.6081008911133,-82.7438888549805,-82.7141571044922,-82.6689834594727,-82.6186828613281,-82.5639343261719,-82.5511322021484,-82.5269012451172,-82.5195159912109,-82.4749221801758,-82.409065246582,-82.3736877441406,-82.3648376464844,-82.370964050293,-82.3228759765625,-82.2581024169922,-82.2746810913086,-82.2787246704102,-82.2582778930664,-82.315559387207,-82.34130859375,-82.345085144043,-82.3580322265625,-82.3508605957031,-82.3601226806641,-82.3713760375977,-82.3896102905273,-82.4379196166992,-82.4667434692383,-82.5246353149414,-82.5700302124023],"lat":[35.1494903564453,35.1849479675293,35.2424659729004,35.267578125,35.2930641174316,35.4180335998535,35.439151763916,35.4551658630371,35.4376754760742,35.4384231567383,35.4268951416016,35.4283218383789,35.4439163208008,35.4444046020508,35.468921661377,35.4573783874512,35.4634590148926,35.4787712097168,35.4951591491699,35.4637298583984,35.4516067504883,35.4356422424316,35.3933563232422,35.3096160888672,35.2854957580566,35.2431907653809,35.2429313659668,35.1926727294922,35.1829490661621,35.1827239990234,35.2082405090332,35.1695594787598,35.1735000610352,35.1545600891113,35.1494903564453]}]],[[{"lng":[-83.6956329345703,-83.7535629272461,-83.791877746582,-83.8368606567383,-83.9216537475586,-83.9608993530273,-83.9922027587891,-84.0038909912109,-84.0308609008789,-84.0292053222656,-84.0063095092773,-84.0126495361328,-83.954704284668,-83.9300765991211,-83.8798904418945,-83.8549880981445,-83.7449569702148,-83.6536102294922,-83.63232421875,-83.5867156982422,-83.5803833007812,-83.5988845825195,-83.6342163085938,-83.6893844604492,-83.6900863647461,-83.6791915893555,-83.6956329345703],"lat":[35.2508087158203,35.2371673583984,35.2486610412598,35.2467498779297,35.2189674377441,35.2153816223145,35.2325019836426,35.2620887756348,35.2925224304199,35.3252906799316,35.372859954834,35.4076232910156,35.4554595947266,35.4490165710449,35.462028503418,35.4501266479492,35.4414024353027,35.4210968017578,35.4367408752441,35.4276885986328,35.4010848999023,35.3623657226562,35.3340339660645,35.3082427978516,35.2877807617188,35.2722206115723,35.2508087158203]}]],[[{"lng":[-77.8365783691406,-77.8300628662109,-77.8051834106445,-77.7606964111328,-77.6984176635742,-77.6844711303711,-77.6660537719727,-77.6218032836914,-77.558723449707,-77.5371780395508,-77.5349731445312,-77.4946746826172,-77.4738845825195,-77.4439468383789,-77.4014892578125,-77.3861618041992,-77.4131546020508,-77.4502105712891,-77.4485092163086,-77.4296493530273,-77.4740982055664,-77.5283126831055,-77.5107727050781,-77.5330581665039,-77.599494934082,-77.7319259643555,-77.7447891235352,-77.754997253418,-77.7516021728516,-77.7641372680664,-77.8365783691406],"lat":[35.1717414855957,35.3423500061035,35.3645896911621,35.3619384765625,35.3698196411133,35.3459396362305,35.339672088623,35.3664855957031,35.3810882568359,35.4019813537598,35.4178237915039,35.4074440002441,35.4215278625488,35.3545951843262,35.3427696228027,35.3292617797852,35.3312187194824,35.302059173584,35.2856750488281,35.259838104248,35.2271919250488,35.2396965026855,35.1557693481445,35.1448822021484,35.0708694458008,35.0008354187012,35.0024452209473,35.0180816650391,35.0956764221191,35.1286125183105,35.1717414855957]}]],[[{"lng":[-82.8876953125,-83.0072784423828,-83.0438537597656,-83.0520858764648,-82.9820785522461,-82.9905853271484,-82.9787139892578,-82.9268569946289,-82.9191131591797,-82.8332366943359,-82.7438888549805,-82.6081008911133,-82.6106109619141,-82.5962371826172,-82.589225769043,-82.5700302124023,-82.6544952392578,-82.6860504150391,-82.6880340576172,-82.6973571777344,-82.7713470458984,-82.8876953125],"lat":[35.0553703308105,35.0242042541504,35.0400695800781,35.0634765625,35.1275062561035,35.1581764221191,35.1857299804688,35.2279052734375,35.2907676696777,35.3155708312988,35.4180335998535,35.2930641174316,35.267578125,35.2424659729004,35.1849479675293,35.1494903564453,35.119457244873,35.1214637756348,35.0978012084961,35.0912322998047,35.0854225158691,35.0553703308105]}]],[[{"lng":[-81.3228225708008,-81.3114242553711,-81.3194274902344,-81.3523483276367,-81.3647994995117,-81.3532638549805,-81.4074172973633,-81.4450759887695,-80.9567718505859,-80.9829254150391,-80.9377593994141,-80.9262771606445,-80.9796447753906,-81.0140533447266,-81.0015182495117,-81.0072784423828,-81.0239639282227,-81.0493011474609,-81.3228225708008],"lat":[35.1637573242188,35.1879501342773,35.260498046875,35.2751045227051,35.3103713989258,35.3273010253906,35.359806060791,35.4133682250977,35.3974533081055,35.3690948486328,35.3649253845215,35.3486747741699,35.3332672119141,35.2499008178711,35.195987701416,35.1632499694824,35.1490325927734,35.1515312194824,35.1637573242188]}]],[[{"lng":[-82.2101745605469,-82.2783279418945,-82.3207702636719,-82.3508605957031,-82.3580322265625,-82.345085144043,-82.34130859375,-82.315559387207,-82.2582778930664,-82.1551284790039,-82.1221694946289,-82.0841522216797,-82.0515823364258,-81.9645156860352,-81.9714431762695,-82.2101745605469],"lat":[35.1931266784668,35.1950073242188,35.1841888427734,35.1926727294922,35.2429313659668,35.2431907653809,35.2854957580566,35.3096160888672,35.3933563232422,35.397102355957,35.3886184692383,35.33935546875,35.323127746582,35.2474250793457,35.1882820129395,35.1931266784668]}]],[[{"lng":[-83.1062850952148,-83.5130081176758,-83.5272216796875,-83.5532379150391,-83.5620422363281,-83.6435623168945,-83.6448745727539,-83.7178039550781,-83.7395248413086,-83.7130813598633,-83.7223281860352,-83.6966934204102,-83.6956329345703,-83.6791915893555,-83.4732284545898,-83.4246826171875,-83.3318176269531,-83.3106384277344,-83.2849807739258,-83.2639389038086,-83.232780456543,-83.2200393676758,-83.2179336547852,-83.1934814453125,-83.1771469116211,-83.1482009887695,-83.1450576782227,-83.1614990234375,-83.1062850952148],"lat":[35.0002784729004,34.9920234680176,35.0197372436523,35.0357627868652,35.0559425354004,35.127498626709,35.1433601379395,35.1388092041016,35.1458396911621,35.1807518005371,35.1972732543945,35.2144355773926,35.2508087158203,35.2722206115723,35.2916450500488,35.3153190612793,35.3193435668945,35.2599868774414,35.226634979248,35.2217712402344,35.2308082580566,35.222526550293,35.158992767334,35.139217376709,35.1114959716797,35.091381072998,35.0832862854004,35.0592231750488,35.0002784729004]}]],[[{"lng":[-78.11376953125,-78.1137390136719,-78.1567611694336,-78.2568054199219,-78.2746124267578,-78.3059005737305,-78.3198852539062,-78.309700012207,-78.3223419189453,-78.3388671875,-78.3638153076172,-78.3866729736328,-78.4010009765625,-78.4992904663086,-78.5419921875,-78.5934753417969,-78.6342544555664,-78.6664123535156,-78.6581573486328,-78.645637512207,-78.6334838867188,-78.6452255249023,-78.6327133178711,-78.6352844238281,-78.6127395629883,-78.5804138183594,-78.5387420654297,-78.4936218261719,-78.4166793823242,-78.3101272583008,-78.2574005126953,-78.1651763916016,-78.1631927490234,-78.170280456543,-78.1552429199219,-78.1429824829102,-78.1954498291016,-78.1560974121094,-78.1358184814453,-78.11376953125],"lat":[34.7209854125977,34.6991806030273,34.6771545410156,34.5537528991699,34.5812759399414,34.6053581237793,34.6300926208496,34.6358528137207,34.6455726623535,34.6826057434082,34.6920433044434,34.7218894958496,34.7593307495117,34.8551063537598,34.9078712463379,34.9434700012207,34.9843673706055,35.0714683532715,35.093620300293,35.1025581359863,35.1537246704102,35.1897277832031,35.1972999572754,35.2109527587891,35.2438316345215,35.2865753173828,35.3151168823242,35.2596206665039,35.2490768432617,35.2802848815918,35.2209510803223,35.1932182312012,35.1822891235352,35.097484588623,35.0681915283203,34.9013252258301,34.7436256408691,34.7284774780273,34.7113456726074,34.7209854125977]}]],[[{"lng":[-76.943244934082,-76.9442291259766,-76.9898910522461,-76.9723358154297,-76.9666061401367,-76.9474334716797,-76.8976058959961,-76.8505706787109,-76.6402130126953,-76.6145172119141,-76.5912094116211,-76.5409774780273,-76.5071258544922,-76.6400909423828,-76.605224609375,-76.6242218017578,-76.6780090332031,-76.8494644165039,-76.9414901733398,-76.943244934082],"lat":[35.0700340270996,35.099723815918,35.1540451049805,35.1567726135254,35.1879653930664,35.2169914245605,35.2515716552734,35.2171669006348,35.237247467041,35.2729187011719,35.3120803833008,35.3036880493164,35.2487678527832,35.1725196838379,35.1385612487793,35.0643501281738,35.0240859985352,34.9822196960449,35.0274276733398,35.0700340270996]}]],[[{"lng":[-84.2910385131836,-84.225944519043,-84.1797256469727,-84.1016464233398,-84.0427780151367,-84.0308609008789,-84.0038909912109,-83.9922027587891,-83.9608993530273,-83.9216537475586,-83.8368606567383,-83.791877746582,-83.7535629272461,-83.6956329345703,-83.6966934204102,-83.7223281860352,-83.7130813598633,-83.7395248413086,-83.7553939819336,-83.8538818359375,-83.9073181152344,-83.9528503417969,-83.9597702026367,-83.9885482788086,-84.1182403564453,-84.3238525390625,-84.2910385131836],"lat":[35.2105445861816,35.261604309082,35.2409896850586,35.2455520629883,35.272575378418,35.2925224304199,35.2620887756348,35.2325019836426,35.2153816223145,35.2189674377441,35.2467498779297,35.2486610412598,35.2371673583984,35.2508087158203,35.2144355773926,35.1972732543945,35.1807518005371,35.1458396911621,35.148063659668,35.1257934570312,35.0704154968262,35.0484428405762,35.0191230773926,34.9890670776367,34.988224029541,34.9890098571777,35.2105445861816]}]],[[{"lng":[-78.4992904663086,-78.5174407958984,-78.831657409668,-78.8962631225586,-78.9125518798828,-78.9286270141602,-78.948486328125,-79.0336074829102,-79.0943145751953,-79.1068420410156,-79.0971755981445,-79.0958938598633,-78.9125823974609,-78.8860168457031,-78.8745727539062,-78.812385559082,-78.716064453125,-78.6127395629883,-78.6352844238281,-78.6327133178711,-78.6452255249023,-78.6334838867188,-78.645637512207,-78.6581573486328,-78.6664123535156,-78.6342544555664,-78.5934753417969,-78.5419921875,-78.4992904663086],"lat":[34.8551063537598,34.8435287475586,34.8443565368652,34.8309020996094,34.8274230957031,34.8797988891602,34.9026832580566,34.9538230895996,35.0378570556641,35.1192245483398,35.1768074035645,35.1899642944336,35.2224655151367,35.2299346923828,35.2425346374512,35.2587203979492,35.2599792480469,35.2438316345215,35.2109527587891,35.1972999572754,35.1897277832031,35.1537246704102,35.1025581359863,35.093620300293,35.0714683532715,34.9843673706055,34.9434700012207,34.9078712463379,34.8551063537598]}]],[[{"lng":[-77.0490036010742,-77.0574645996094,-77.0731353759766,-77.1006927490234,-77.1598205566406,-77.1642608642578,-77.1928405761719,-77.180061340332,-77.2062683105469,-77.2229919433594,-77.2854232788086,-77.3351364135742,-77.3699722290039,-77.4996337890625,-77.6392517089844,-77.682991027832,-77.7319259643555,-77.599494934082,-77.5330581665039,-77.5107727050781,-77.5283126831055,-77.4740982055664,-77.4234085083008,-77.2055130004883,-77.1792449951172,-77.1404495239258,-77.1311187744141,-77.1105880737305,-77.1033706665039,-77.1095352172852,-77.0490036010742],"lat":[34.8311920166016,34.8150482177734,34.8154411315918,34.7974967956543,34.7884979248047,34.7745170593262,34.8042869567871,34.8321495056152,34.8355026245117,34.8845100402832,34.9264030456543,34.9325408935547,34.914249420166,34.9625053405762,34.9840202331543,34.966724395752,35.0008354187012,35.0708694458008,35.1448822021484,35.1557693481445,35.2396965026855,35.2271919250488,35.1943054199219,35.0880546569824,35.0533599853516,35.0542373657227,35.0692825317383,35.0660362243652,35.0242614746094,34.8304290771484,34.8311920166016]}]],[[{"lng":[-80.8401641845703,-80.7972183227539,-80.7596817016602,-80.5396423339844,-80.5029449462891,-80.4832305908203,-80.4532852172852,-80.428581237793,-80.3987503051758,-80.3729476928711,-80.3487243652344,-80.3256759643555,-80.2751235961914,-80.3252792358398,-80.5615463256836,-80.8000640869141,-80.7856521606445,-80.8401641845703],"lat":[35.0020179748535,35.0281982421875,35.03662109375,35.2058181762695,35.1869125366211,35.1829109191895,35.1607856750488,35.1685943603516,35.1646118164062,35.177864074707,35.1706733703613,35.1739158630371,35.1931114196777,34.8147621154785,34.8152275085449,34.8161125183105,34.9406394958496,35.0020179748535]}]],[[{"lng":[-79.9199523925781,-80.3252792358398,-80.2751235961914,-80.2610931396484,-80.2475891113281,-80.2198867797852,-80.1901016235352,-80.1681289672852,-80.1591415405273,-80.1546859741211,-80.1316146850586,-80.1102752685547,-80.0714111328125,-80.0651779174805,-80.0584487915039,-80.0028076171875,-79.9752807617188,-79.9449157714844,-79.9274978637695,-79.9151382446289,-79.9084320068359,-79.882080078125,-79.8939208984375,-79.8670501708984,-79.8537063598633,-79.9014205932617,-79.9199523925781],"lat":[34.807918548584,34.8147621154785,35.1931114196777,35.2040405273438,35.2045249938965,35.1591835021973,35.1678581237793,35.149730682373,35.1547393798828,35.176082611084,35.1711158752441,35.1938362121582,35.1507415771484,35.1365814208984,35.1039848327637,35.0899276733398,35.0967330932617,35.1153373718262,35.115779876709,35.1066932678223,35.0789947509766,35.0444641113281,35.0072441101074,34.967716217041,34.9045829772949,34.852409362793,34.807918548584]}]],[[{"lng":[-79.3403015136719,-79.3606796264648,-79.3984146118164,-79.4553604125977,-79.3500518798828,-79.2352905273438,-79.1472778320312,-79.1220016479492,-79.0971755981445,-79.1068420410156,-79.0943145751953,-79.0336074829102,-79.1818542480469,-79.3403015136719],"lat":[34.8356094360352,34.944709777832,34.9962158203125,35.0373573303223,35.140796661377,35.2032089233398,35.1735496520996,35.1701850891113,35.1768074035645,35.1192245483398,35.0378570556641,34.9538230895996,34.8318824768066,34.8356094360352]}]],[[{"lng":[-76.5189437866211,-76.5395965576172,-76.5858764648438,-76.6079483032227,-76.6089172363281,-76.6338195800781,-76.4084320068359,-76.2734451293945,-76.2976303100586,-76.232795715332,-76.2102355957031,-76.1673049926758,-76.1047439575195,-76.0212097167969,-75.9080276489258,-75.8914947509766,-76.052978515625,-76.1816177368164,-76.4988021850586,-76.5337524414062,-76.4584350585938,-76.4465255737305,-76.5189437866211],"lat":[35.5776443481445,35.5940361022949,35.6094551086426,35.635066986084,35.6641540527344,35.7030029296875,35.6991157531738,35.6894989013672,35.6116943359375,35.5946655273438,35.6043891906738,35.6968421936035,35.6636390686035,35.6690940856934,35.6656379699707,35.6312675476074,35.4146041870117,35.3415336608887,35.4162178039551,35.4501991271973,35.5044250488281,35.5508651733398,35.5776443481445]}],[{"lng":[-76.0170059204102,-75.9756927490234,-75.8539352416992,-75.7637023925781,-75.8115386962891,-75.9039459228516,-76.0028762817383,-76.0170059204102],"lat":[35.0694351196289,35.1161842346191,35.1672821044922,35.1922836303711,35.1640205383301,35.1324920654297,35.0694351196289,35.0694351196289]}]],[[{"lng":[-77.6898345947266,-77.9266738891602,-77.9393081665039,-77.9891510009766,-78.0173645019531,-78.11376953125,-78.1358184814453,-78.1560974121094,-78.1954498291016,-78.1429824829102,-78.1552429199219,-78.170280456543,-78.1631927490234,-78.087532043457,-78.036506652832,-78.0021591186523,-77.9639282226562,-77.944694519043,-77.9139785766602,-77.894157409668,-77.8876724243164,-77.8365783691406,-77.7641372680664,-77.7516021728516,-77.754997253418,-77.7447891235352,-77.7319259643555,-77.682991027832,-77.6552658081055,-77.6898345947266],"lat":[34.7201957702637,34.7110061645508,34.7194061279297,34.7170944213867,34.728931427002,34.7209854125977,34.7113456726074,34.7284774780273,34.7436256408691,34.9013252258301,35.0681915283203,35.097484588623,35.1822891235352,35.1701850891113,35.1856918334961,35.1864776611328,35.1640243530273,35.1682319641113,35.1599731445312,35.1441802978516,35.1549644470215,35.1717414855957,35.1286125183105,35.0956764221191,35.0180816650391,35.0024452209473,35.0008354187012,34.966724395752,34.9189338684082,34.7201957702637]}]],[[{"lng":[-79.6859588623047,-79.9199523925781,-79.9014205932617,-79.8537063598633,-79.8670501708984,-79.8939208984375,-79.882080078125,-79.9084320068359,-79.9151382446289,-79.9274978637695,-79.9449157714844,-79.9752807617188,-80.0028076171875,-80.0584487915039,-80.0651779174805,-80.0106811523438,-79.9758224487305,-79.9094619750977,-79.8363189697266,-79.7598419189453,-79.6979522705078,-79.6372985839844,-79.6074676513672,-79.5756301879883,-79.5741958618164,-79.5529022216797,-79.4950637817383,-79.4553604125977,-79.5471038818359,-79.5730438232422,-79.5805206298828,-79.5655899047852,-79.6219253540039,-79.611442565918,-79.6601486206055,-79.6859588623047],"lat":[34.8052558898926,34.807918548584,34.852409362793,34.9045829772949,34.967716217041,35.0072441101074,35.0444641113281,35.0789947509766,35.1066932678223,35.115779876709,35.1153373718262,35.0967330932617,35.0899276733398,35.1039848327637,35.1365814208984,35.1371421813965,35.1507568359375,35.1557197570801,35.1738014221191,35.1673126220703,35.1730728149414,35.1538352966309,35.1591873168945,35.1191253662109,35.0732650756836,35.0641059875488,35.0625076293945,35.0373573303223,35.0000610351562,34.9783668518066,34.9447898864746,34.9075012207031,34.8495712280273,34.8177490234375,34.8178977966309,34.8052558898926]}]],[[{"lng":[-83.9379959106445,-83.9885482788086,-83.9597702026367,-83.9528503417969,-83.9073181152344,-83.8538818359375,-83.7553939819336,-83.7395248413086,-83.7178039550781,-83.6448745727539,-83.6435623168945,-83.5620422363281,-83.5532379150391,-83.5272216796875,-83.5130081176758,-83.5494155883789,-83.9379959106445],"lat":[34.9893913269043,34.9890670776367,35.0191230773926,35.0484428405762,35.0704154968262,35.1257934570312,35.148063659668,35.1458396911621,35.1388092041016,35.1433601379395,35.127498626709,35.0559425354004,35.0357627868652,35.0197372436523,34.9920234680176,34.9895362854004,34.9893913269043]}]],[[{"lng":[-76.8976058959961,-76.9474334716797,-76.9666061401367,-76.9723358154297,-76.9898910522461,-76.9442291259766,-76.943244934082,-77.0478820800781,-77.1046752929688,-77.1105880737305,-77.1311187744141,-77.1404495239258,-77.1792449951172,-77.2055130004883,-77.4234085083008,-77.4740982055664,-77.4296493530273,-77.4485092163086,-77.4502105712891,-77.4131546020508,-77.3861618041992,-77.3542175292969,-77.2938003540039,-77.2641906738281,-77.2443008422852,-77.2405471801758,-77.2112045288086,-77.1949615478516,-76.8976058959961],"lat":[35.2515716552734,35.2169914245605,35.1879653930664,35.1567726135254,35.1540451049805,35.099723815918,35.0700340270996,35.092155456543,35.0879859924316,35.0660362243652,35.0692825317383,35.0542373657227,35.0533599853516,35.0880546569824,35.1943054199219,35.2271919250488,35.259838104248,35.2856750488281,35.302059173584,35.3312187194824,35.3292617797852,35.3235549926758,35.373950958252,35.3501129150391,35.354190826416,35.3799858093262,35.3956413269043,35.4229545593262,35.2515716552734]}],[{"lng":[-76.6470489501953,-76.625617980957,-76.7502136230469,-77.0490036010742,-77.1095352172852,-77.1033706665039,-77.1105880737305,-77.0011901855469,-76.9131088256836,-76.6470489501953],"lat":[34.9063339233398,34.8906478881836,34.848274230957,34.8311920166016,34.8304290771484,35.0242614746094,35.0660362243652,35.0528869628906,34.9364700317383,34.9063339233398]}]],[[{"lng":[-79.4559707641602,-79.6675033569336,-79.6859588623047,-79.6601486206055,-79.611442565918,-79.6219253540039,-79.5655899047852,-79.5805206298828,-79.5730438232422,-79.5471038818359,-79.4553604125977,-79.3984146118164,-79.3606796264648,-79.3403015136719,-79.3444519042969,-79.3279724121094,-79.4559707641602],"lat":[34.634090423584,34.8006629943848,34.8052558898926,34.8178977966309,34.8177490234375,34.8495712280273,34.9075012207031,34.9447898864746,34.9783668518066,35.0000610351562,35.0373573303223,34.9962158203125,34.944709777832,34.8356094360352,34.808837890625,34.7724113464355,34.634090423584]}]],[[{"lng":[-77.5386352539062,-77.5763320922852,-77.6898345947266,-77.6552658081055,-77.682991027832,-77.6392517089844,-77.4996337890625,-77.3699722290039,-77.3351364135742,-77.2854232788086,-77.2229919433594,-77.2062683105469,-77.180061340332,-77.1928405761719,-77.1642608642578,-77.1489562988281,-77.12939453125,-77.1566009521484,-77.5386352539062],"lat":[34.4570083618164,34.4693450927734,34.7201957702637,34.9189338684082,34.966724395752,34.9840202331543,34.9625053405762,34.914249420166,34.9325408935547,34.9264030456543,34.8845100402832,34.8355026245117,34.8321495056152,34.8042869567871,34.7745170593262,34.764331817627,34.6849136352539,34.6606369018555,34.4570083618164]}]],[[{"lng":[-78.8645095825195,-78.9194717407227,-78.9507369995117,-78.9753646850586,-79.0022354125977,-79.0064163208008,-79.0294723510742,-79.0409469604492,-79.0745010375977,-79.4471435546875,-79.4559707641602,-79.3279724121094,-79.3444519042969,-79.3403015136719,-79.1818542480469,-79.0336074829102,-78.948486328125,-78.9286270141602,-78.9125518798828,-78.8962631225586,-78.8417053222656,-78.8033218383789,-78.8526000976562,-78.8456649780273,-78.8645095825195],"lat":[34.4771957397461,34.4536399841309,34.4493827819824,34.3991661071777,34.3880424499512,34.366268157959,34.3473739624023,34.3192977905273,34.304573059082,34.6190605163574,34.634090423584,34.7724113464355,34.808837890625,34.8356094360352,34.8318824768066,34.9538230895996,34.9026832580566,34.8797988891602,34.8274230957031,34.8309020996094,34.7350006103516,34.6851043701172,34.5665817260742,34.507453918457,34.4771957397461]}]],[[{"lng":[-77.1489562988281,-77.1642608642578,-77.1598205566406,-77.1006927490234,-77.0731353759766,-77.0574645996094,-77.0490036010742,-76.7502136230469,-76.625617980957,-76.6470489501953,-76.6700592041016,-76.5297775268555,-76.4450302124023,-76.4240570068359,-76.364013671875,-76.3294067382812,-76.3147125244141,-76.4699020385742,-76.6253128051758,-77.0505294799805,-77.1489562988281],"lat":[34.764331817627,34.7745170593262,34.7884979248047,34.7974967956543,34.8154411315918,34.8150482177734,34.8311920166016,34.848274230957,34.8906478881836,34.9063339233398,34.9699974060059,34.9746360778809,35.016658782959,34.9462471008301,34.9431114196777,34.9759559631348,34.9488143920898,34.7850608825684,34.7197532653809,34.698917388916,34.764331817627]}],[{"lng":[-76.5442733764648,-76.5551452636719,-76.5377502441406,-76.4844818115234,-76.4322967529297,-76.3746795654297,-76.2877044677734,-76.4692611694336,-76.5442733764648],"lat":[34.5878295898438,34.6106605529785,34.6139221191406,34.6976318359375,34.760684967041,34.8128700256348,34.877010345459,34.6932830810547,34.5878295898438]}]],[[{"lng":[-78.2614974975586,-78.3289794921875,-78.4379425048828,-78.6380844116211,-78.6598205566406,-78.6684951782227,-78.8118896484375,-78.8363571166992,-78.8645095825195,-78.8456649780273,-78.8526000976562,-78.8033218383789,-78.8417053222656,-78.8962631225586,-78.831657409668,-78.5174407958984,-78.4992904663086,-78.4010009765625,-78.3866729736328,-78.3638153076172,-78.3388671875,-78.3223419189453,-78.309700012207,-78.3198852539062,-78.3059005737305,-78.2746124267578,-78.2568054199219,-78.2624053955078,-78.2550582885742,-78.169677734375,-78.2614974975586],"lat":[34.3947868347168,34.3644218444824,34.3718414306641,34.4511451721191,34.4509429931641,34.4651298522949,34.4553184509277,34.4591979980469,34.4771957397461,34.507453918457,34.5665817260742,34.6851043701172,34.7350006103516,34.8309020996094,34.8443565368652,34.8435287475586,34.8551063537598,34.7593307495117,34.7218894958496,34.6920433044434,34.6826057434082,34.6455726623535,34.6358528137207,34.6300926208496,34.6053581237793,34.5812759399414,34.5537528991699,34.5279388427734,34.507381439209,34.4624519348145,34.3947868347168]}]],[[{"lng":[-78.0259246826172,-78.130241394043,-78.15478515625,-78.1846160888672,-78.1945648193359,-78.2614974975586,-78.169677734375,-78.2550582885742,-78.2624053955078,-78.2568054199219,-78.1567611694336,-78.1137390136719,-78.11376953125,-78.0173645019531,-77.9891510009766,-77.9393081665039,-77.9266738891602,-77.6898345947266,-77.5763320922852,-77.5386352539062,-77.5867614746094,-77.6097259521484,-77.6026153564453,-77.7505264282227,-77.8091430664062,-77.8283843994141,-77.888069152832,-77.9217834472656,-77.9439392089844,-77.9498138427734,-77.9790725708008,-77.9944534301758,-77.9866790771484,-78.0259246826172],"lat":[34.3287696838379,34.3641242980957,34.3622436523438,34.3481712341309,34.3510513305664,34.3947868347168,34.4624519348145,34.507381439209,34.5279388427734,34.5537528991699,34.6771545410156,34.6991806030273,34.7209854125977,34.728931427002,34.7170944213867,34.7194061279297,34.7110061645508,34.7201957702637,34.4693450927734,34.4570083618164,34.4209365844727,34.4350433349609,34.4126281738281,34.305046081543,34.359432220459,34.3879699707031,34.364070892334,34.3733139038086,34.3564376831055,34.3660850524902,34.3756866455078,34.3623161315918,34.339916229248,34.3287696838379]}]],[[{"lng":[-78.6557159423828,-79.0745010375977,-79.0409469604492,-79.0294723510742,-79.0064163208008,-79.0022354125977,-78.9753646850586,-78.9507369995117,-78.9194717407227,-78.8645095825195,-78.8363571166992,-78.8118896484375,-78.6684951782227,-78.6598205566406,-78.6380844116211,-78.4379425048828,-78.3289794921875,-78.2614974975586,-78.1945648193359,-78.1846160888672,-78.15478515625,-78.2610626220703,-78.3735733032227,-78.3611221313477,-78.4254302978516,-78.4927444458008,-78.5272369384766,-78.5442810058594,-78.5634307861328,-78.5877838134766,-78.6302719116211,-78.6347198486328,-78.6557159423828],"lat":[33.948673248291,34.304573059082,34.3192977905273,34.3473739624023,34.366268157959,34.3880424499512,34.3991661071777,34.4493827819824,34.4536399841309,34.4771957397461,34.4591979980469,34.4553184509277,34.4651298522949,34.4509429931641,34.4511451721191,34.3718414306641,34.3644218444824,34.3947868347168,34.3510513305664,34.3481712341309,34.3622436523438,34.2152633666992,34.2023506164551,34.1867218017578,34.1380653381348,34.158504486084,34.154857635498,34.134162902832,34.0589447021484,34.0306053161621,34.0102005004883,33.9779777526855,33.948673248291]}]],[[{"lng":[-77.9607315063477,-77.9658660888672,-77.9752807617188,-77.9831466674805,-78.0002212524414,-77.9953918457031,-78.0070190429688,-78.0113067626953,-78.0259246826172,-77.9866790771484,-77.9944534301758,-77.9790725708008,-77.9498138427734,-77.9439392089844,-77.9217834472656,-77.888069152832,-77.8283843994141,-77.8091430664062,-77.7505264282227,-77.864387512207,-77.894401550293,-77.9267578125,-77.9607315063477],"lat":[34.1892433166504,34.2422866821289,34.2433624267578,34.2616806030273,34.2678833007812,34.2827987670898,34.2848167419434,34.312614440918,34.3287696838379,34.339916229248,34.3623161315918,34.3756866455078,34.3660850524902,34.3564376831055,34.3733139038086,34.364070892334,34.3879699707031,34.359432220459,34.305046081543,34.1927375793457,34.0691795349121,34.0620346069336,34.1892433166504]}]],[[{"lng":[-78.6557159423828,-78.6347198486328,-78.6302719116211,-78.5877838134766,-78.5634307861328,-78.5442810058594,-78.5272369384766,-78.4927444458008,-78.4254302978516,-78.3611221313477,-78.3735733032227,-78.2610626220703,-78.15478515625,-78.130241394043,-78.0259246826172,-78.0113067626953,-78.0070190429688,-77.9953918457031,-78.0002212524414,-77.9831466674805,-77.9752807617188,-77.9658660888672,-77.9607315063477,-77.9585266113281,-78.0348052978516,-78.579719543457,-78.6557159423828],"lat":[33.948673248291,33.9779777526855,34.0102005004883,34.0306053161621,34.0589447021484,34.134162902832,34.154857635498,34.158504486084,34.1380653381348,34.1867218017578,34.2023506164551,34.2152633666992,34.3622436523438,34.3641242980957,34.3287696838379,34.312614440918,34.2848167419434,34.2827987670898,34.2678833007812,34.2616806030273,34.2433624267578,34.2422866821289,34.1892433166504,33.9925804138184,33.9142913818359,33.8819923400879,33.948673248291]}]]],null,null,{"interactive":true,"className":"","stroke":true,"color":"grey","weight":5,"opacity":0.5,"fill":true,"fillColor":"grey","fillOpacity":0.2,"smoothFactor":1,"noClip":false},null,null,null,{"interactive":false,"permanent":false,"direction":"auto","opacity":1,"offset":[0,0],"textsize":"10px","textOnly":false,"className":"","sticky":true},null]},{"method":"addPolygons","args":[[[[{"lng":[-81.4727554321289,-81.5408401489258,-81.5619812011719,-81.6330642700195,-81.7410736083984,-81.6982803344727,-81.7027969360352,-81.6699981689453,-81.3452987670898,-81.347541809082,-81.3247756958008,-81.3133239746094,-81.2662353515625,-81.2628402709961,-81.2406921386719,-81.2398910522461,-81.2642440795898,-81.3289947509766,-81.3613739013672,-81.3656921386719,-81.354133605957,-81.3674545288086,-81.4063873291016,-81.4123306274414,-81.431037902832,-81.4528884887695,-81.4727554321289],"lat":[36.2343559265137,36.2725067138672,36.2735939025879,36.3406867980957,36.3917846679688,36.4717788696289,36.5193405151367,36.5896492004395,36.5728645324707,36.537914276123,36.5136795043945,36.4806976318359,36.4372062683105,36.4050407409668,36.3794174194336,36.365364074707,36.3524131774902,36.3635025024414,36.3531608581543,36.3390502929688,36.2997169494629,36.2786979675293,36.2850532531738,36.2672920227051,36.2607192993164,36.2395858764648,36.2343559265137]}]],[[{"lng":[-81.2398910522461,-81.2406921386719,-81.2628402709961,-81.2662353515625,-81.3133239746094,-81.3247756958008,-81.347541809082,-81.3452987670898,-80.9034423828125,-80.9335479736328,-80.9657745361328,-80.9496688842773,-80.9563903808594,-80.9779510498047,-80.9828414916992,-81.0027770996094,-81.0246429443359,-81.0428009033203,-81.0842514038086,-81.0985641479492,-81.1133117675781,-81.1293792724609,-81.1383972167969,-81.1533660888672,-81.1766738891602,-81.2398910522461],"lat":[36.365364074707,36.3794174194336,36.4050407409668,36.4372062683105,36.4806976318359,36.5136795043945,36.537914276123,36.5728645324707,36.5652122497559,36.4983139038086,36.4672203063965,36.4147338867188,36.4037971496582,36.3913764953613,36.3718338012695,36.3666801452637,36.3778343200684,36.4103355407715,36.4299201965332,36.43115234375,36.4228515625,36.4263305664062,36.4176254272461,36.4247398376465,36.4154434204102,36.365364074707]}]],[[{"lng":[-80.4563446044922,-80.4763870239258,-80.5368804931641,-80.5450134277344,-80.5541534423828,-80.5905990600586,-80.6243133544922,-80.6674423217773,-80.6966552734375,-80.7240371704102,-80.7343673706055,-80.7525634765625,-80.7662963867188,-80.7826995849609,-80.874382019043,-80.8708648681641,-80.8889236450195,-80.9245681762695,-80.9563903808594,-80.9496688842773,-80.9657745361328,-80.9335479736328,-80.9034423828125,-80.8381576538086,-80.6110534667969,-80.4353103637695,-80.453010559082,-80.4563446044922],"lat":[36.2425575256348,36.2547264099121,36.2567367553711,36.2766571044922,36.2784309387207,36.2682762145996,36.2730979919434,36.2461013793945,36.259090423584,36.258472442627,36.2647590637207,36.2582969665527,36.2618370056152,36.2485771179199,36.2338829040527,36.3246231079102,36.3544273376465,36.3727531433105,36.4037971496582,36.4147338867188,36.4672203063965,36.4983139038086,36.5652122497559,36.5634384155273,36.5572967529297,36.5510444641113,36.2570877075195,36.2425575256348]}]],[[{"lng":[-76.0089721679688,-76.0173492431641,-76.0328750610352,-76.0439529418945,-76.095085144043,-76.1609268188477,-76.1581497192383,-76.1682891845703,-76.3302536010742,-76.1273956298828,-76.0459594726562,-76.0332107543945,-76.091064453125,-75.97607421875,-75.9697647094727,-76.0016098022461,-75.9512557983398,-75.9281234741211,-75.9245910644531,-75.8000564575195,-75.7988510131836,-75.8551635742188,-75.9137649536133,-75.9575119018555,-75.9419326782227,-76.0089721679688],"lat":[36.3195953369141,36.3377304077148,36.3359756469727,36.3535919189453,36.3489151000977,36.3918991088867,36.4126892089844,36.4270858764648,36.5560569763184,36.5571632385254,36.5569534301758,36.5143737792969,36.5035667419434,36.4362144470215,36.4151191711426,36.4189147949219,36.3654708862305,36.4232444763184,36.3509483337402,36.1128158569336,36.0728187561035,36.1056671142578,36.244800567627,36.2594528198242,36.2943382263184,36.3195953369141]}],[{"lng":[-76.0271682739258,-75.998664855957,-75.9119186401367,-75.9248046875,-75.9772796630859,-75.9762878417969,-76.0271682739258],"lat":[36.5567169189453,36.5566520690918,36.5425300598145,36.4739761352539,36.4780158996582,36.5179252624512,36.5567169189453]}],[{"lng":[-75.901985168457,-75.8781661987305,-75.7731552124023,-75.7831726074219,-75.901985168457],"lat":[36.5561981201172,36.5558738708496,36.2292556762695,36.2251930236816,36.5561981201172]}]],[[{"lng":[-77.2176666259766,-77.2346115112305,-77.2986145019531,-77.2935104370117,-77.3094787597656,-77.3349914550781,-77.389030456543,-77.3786239624023,-77.4134674072266,-77.4188537597656,-77.4541168212891,-77.5380783081055,-77.5574340820312,-77.5719528198242,-77.580078125,-77.559684753418,-77.560417175293,-77.6359710693359,-77.650993347168,-77.6988754272461,-77.7494049072266,-77.9012069702148,-77.8988571166992,-77.7639312744141,-77.3200531005859,-77.1773529052734,-77.1544189453125,-77.0921325683594,-77.075309753418,-77.0832748413086,-77.127326965332,-77.1393203735352,-77.141960144043,-77.2176666259766],"lat":[36.2409820556641,36.214599609375,36.2115287780762,36.1628608703613,36.1627655029297,36.1828498840332,36.2031021118164,36.2400856018066,36.2558174133301,36.281795501709,36.3197555541992,36.3024597167969,36.3042144775391,36.3144950866699,36.3282699584961,36.3759460449219,36.4063568115234,36.4405364990234,36.4725875854492,36.4789581298828,36.4735717773438,36.5098876953125,36.5529441833496,36.5534400939941,36.5539169311523,36.5562858581543,36.526252746582,36.5075187683105,36.4835166931152,36.4700965881348,36.4707107543945,36.4564781188965,36.417064666748,36.2409820556641]}]],[[{"lng":[-76.7450637817383,-76.9806900024414,-76.9947509765625,-77.1300735473633,-77.2176666259766,-77.141960144043,-77.1393203735352,-77.127326965332,-77.0832748413086,-77.075309753418,-77.0921325683594,-77.1544189453125,-77.1773529052734,-76.9241333007812,-76.9084243774414,-76.9457702636719,-76.9536743164062,-76.9435119628906,-76.9240798950195,-76.7413482666016,-76.7074966430664,-76.7450637817383],"lat":[36.2339172363281,36.2302360534668,36.2355804443359,36.2334632873535,36.2409820556641,36.417064666748,36.4564781188965,36.4707107543945,36.4700965881348,36.4835166931152,36.5075187683105,36.526252746582,36.5562858581543,36.5541458129883,36.5042839050293,36.4589614868164,36.4192314147949,36.4017295837402,36.3924446105957,36.3151664733887,36.2661323547363,36.2339172363281]}]],[[{"lng":[-76.0089721679688,-75.9571838378906,-75.9813385009766,-76.1831665039062,-76.1934814453125,-76.216194152832,-76.2385330200195,-76.2612838745117,-76.2741317749023,-76.3033599853516,-76.3213653564453,-76.4059677124023,-76.4983444213867,-76.5635833740234,-76.49755859375,-76.3302536010742,-76.1682891845703,-76.1581497192383,-76.1609268188477,-76.095085144043,-76.0439529418945,-76.0328750610352,-76.0173492431641,-76.0089721679688],"lat":[36.3195953369141,36.1937713623047,36.1697273254395,36.3152389526367,36.3248519897461,36.3278465270996,36.3612289428711,36.3637580871582,36.3814086914062,36.391845703125,36.4096450805664,36.4471588134766,36.5039024353027,36.5552520751953,36.5558128356934,36.5560569763184,36.4270858764648,36.4126892089844,36.3918991088867,36.3489151000977,36.3535919189453,36.3359756469727,36.3377304077148,36.3195953369141]}]],[[{"lng":[-76.5625076293945,-76.6042404174805,-76.6482162475586,-76.6887359619141,-76.7766418457031,-76.9240798950195,-76.9435119628906,-76.9536743164062,-76.9457702636719,-76.9084243774414,-76.9241333007812,-76.921630859375,-76.5635833740234,-76.4983444213867,-76.5024642944336,-76.4603500366211,-76.5625076293945],"lat":[36.3405685424805,36.3149833679199,36.315315246582,36.2945175170898,36.3583297729492,36.3924446105957,36.4017295837402,36.4192314147949,36.4589614868164,36.5042839050293,36.5541458129883,36.5541572570801,36.5552520751953,36.5039024353027,36.4522857666016,36.3738975524902,36.3405685424805]}]],[[{"lng":[-78.3087615966797,-78.2829284667969,-78.3212509155273,-78.0516662597656,-77.8988571166992,-77.9012069702148,-77.9169235229492,-77.9301376342773,-77.9520874023438,-78.0062866210938,-78.058349609375,-78.1096267700195,-78.1347198486328,-78.3087615966797],"lat":[36.2600402832031,36.2918815612793,36.5455322265625,36.5524749755859,36.5529441833496,36.5098876953125,36.5006370544434,36.3529853820801,36.2812309265137,36.1959457397461,36.2113227844238,36.213508605957,36.2365837097168,36.2600402832031]}]],[[{"lng":[-80.0256729125977,-80.453010559082,-80.4353103637695,-80.048095703125,-80.024055480957,-80.0256729125977],"lat":[36.2502326965332,36.2570877075195,36.5510444641113,36.5471343994141,36.5450248718262,36.2502326965332]}]],[[{"lng":[-79.5305099487305,-79.5102996826172,-79.2170639038086,-79.1443252563477,-79.1592712402344,-79.2584991455078,-79.5305099487305],"lat":[36.2461357116699,36.5476570129395,36.5497817993164,36.5460586547852,36.2336692810059,36.2356872558594,36.2461357116699]}]],[[{"lng":[-79.5305099487305,-79.5305786132812,-80.0256729125977,-80.024055480957,-79.7174453735352,-79.5102996826172,-79.5305099487305],"lat":[36.2461357116699,36.236156463623,36.2502326965332,36.5450248718262,36.5478897094727,36.5476570129395,36.2461357116699]}]],[[{"lng":[-78.7491226196289,-78.7884140014648,-78.8040542602539,-78.8103561401367,-78.8068008422852,-78.7966995239258,-78.7373886108398,-78.4588088989258,-78.463752746582,-78.5025024414062,-78.51708984375,-78.5147171020508,-78.4925231933594,-78.5458526611328,-78.5480270385742,-78.6955718994141,-78.7491226196289],"lat":[36.063591003418,36.062183380127,36.080940246582,36.114574432373,36.231575012207,36.5435333251953,36.5460739135742,36.5414810180664,36.5238571166992,36.5043907165527,36.461483001709,36.1752243041992,36.1735877990723,36.0680885314941,36.0141258239746,36.0666465759277,36.063591003418]}]],[[{"lng":[-78.8068008422852,-78.9510803222656,-79.1592712402344,-79.1443252563477,-78.7966995239258,-78.8068008422852],"lat":[36.231575012207,36.2338371276855,36.2336692810059,36.5460586547852,36.5435333251953,36.231575012207]}]],[[{"lng":[-78.4925231933594,-78.5147171020508,-78.51708984375,-78.5025024414062,-78.463752746582,-78.4588088989258,-78.3212509155273,-78.2829284667969,-78.3087615966797,-78.3460464477539,-78.3808517456055,-78.4169540405273,-78.4925231933594],"lat":[36.1735877990723,36.1752243041992,36.461483001709,36.5043907165527,36.5238571166992,36.5414810180664,36.5455322265625,36.2918815612793,36.2600402832031,36.2251815795898,36.2247505187988,36.1621742248535,36.1735877990723]}]],[[{"lng":[-77.3322067260742,-77.4053115844727,-77.4257431030273,-77.4382171630859,-77.4639739990234,-77.5251312255859,-77.5358581542969,-77.5366897583008,-77.5478820800781,-77.6062545776367,-77.6423568725586,-77.6856231689453,-77.7005081176758,-77.7201843261719,-77.7471694946289,-77.8013687133789,-77.8113021850586,-77.8868408203125,-77.917839050293,-77.9224014282227,-77.9392318725586,-77.9558639526367,-77.9733963012695,-77.9851150512695,-78.0062866210938,-77.9520874023438,-77.9301376342773,-77.9169235229492,-77.9012069702148,-77.7494049072266,-77.6988754272461,-77.650993347168,-77.6359710693359,-77.560417175293,-77.559684753418,-77.580078125,-77.5719528198242,-77.5574340820312,-77.5380783081055,-77.4541168212891,-77.4188537597656,-77.4134674072266,-77.3786239624023,-77.389030456543,-77.3349914550781,-77.3094787597656,-77.2935104370117,-77.2704086303711,-77.2559127807617,-77.2574996948242,-77.2410354614258,-77.3098754882812,-77.3322067260742],"lat":[36.0679817199707,35.9947166442871,35.9960632324219,36.0140342712402,36.0263824462891,36.0353851318359,36.0555725097656,36.0823631286621,36.088493347168,36.0973930358887,36.1267013549805,36.1466026306152,36.1441688537598,36.1341133117676,36.1464233398438,36.1442565917969,36.1358184814453,36.1444396972656,36.1567802429199,36.1777305603027,36.1875495910645,36.1837501525879,36.1890411376953,36.1774444580078,36.1959457397461,36.2812309265137,36.3529853820801,36.5006370544434,36.5098876953125,36.4735717773438,36.4789581298828,36.4725875854492,36.4405364990234,36.4063568115234,36.3759460449219,36.3282699584961,36.3144950866699,36.3042144775391,36.3024597167969,36.3197555541992,36.281795501709,36.2558174133301,36.2400856018066,36.2031021118164,36.1828498840332,36.1627655029297,36.1628608703613,36.1555252075195,36.130687713623,36.1184730529785,36.1013031005859,36.0874443054199,36.0679817199707]}]],[[{"lng":[-76.2989273071289,-76.3242340087891,-76.3724212646484,-76.4603500366211,-76.5024642944336,-76.4983444213867,-76.4059677124023,-76.3213653564453,-76.3033599853516,-76.2741317749023,-76.2612838745117,-76.2385330200195,-76.216194152832,-76.1934814453125,-76.1831665039062,-76.2189025878906,-76.1127090454102,-76.1419372558594,-76.234977722168,-76.2989273071289],"lat":[36.2142295837402,36.2336235046387,36.2523460388184,36.3738975524902,36.4522857666016,36.5039024353027,36.4471588134766,36.4096450805664,36.391845703125,36.3814086914062,36.3637580871582,36.3612289428711,36.3278465270996,36.3248519897461,36.3152389526367,36.2966079711914,36.1744194030762,36.1476898193359,36.1633605957031,36.2142295837402]}]],[[{"lng":[-81.0205688476562,-81.0840835571289,-81.1240615844727,-81.1574630737305,-81.2360076904297,-81.3218688964844,-81.3473510742188,-81.3887100219727,-81.3981475830078,-81.4296340942383,-81.4544372558594,-81.5173110961914,-81.5461044311523,-81.4993209838867,-81.4508056640625,-81.4727554321289,-81.4528884887695,-81.431037902832,-81.4123306274414,-81.4063873291016,-81.3674545288086,-81.354133605957,-81.3656921386719,-81.3613739013672,-81.3289947509766,-81.2642440795898,-81.2398910522461,-81.1766738891602,-81.1533660888672,-81.1383972167969,-81.1293792724609,-81.1133117675781,-81.0985641479492,-81.0842514038086,-81.0428009033203,-81.0246429443359,-81.0027770996094,-80.9828414916992,-80.9779510498047,-80.9563903808594,-80.9245681762695,-80.8889236450195,-80.8708648681641,-80.874382019043,-80.8774108886719,-80.9889526367188,-81.0205688476562],"lat":[36.0349349975586,36.0207672119141,36.0312843322754,36.0209808349609,36.0238227844238,35.9893264770508,36.0153579711914,36.0375671386719,36.0560569763184,36.0656623840332,36.0830574035645,36.0954322814941,36.1113929748535,36.1360359191895,36.1906204223633,36.2343559265137,36.2395858764648,36.2607192993164,36.2672920227051,36.2850532531738,36.2786979675293,36.2997169494629,36.3390502929688,36.3531608581543,36.3635025024414,36.3524131774902,36.365364074707,36.4154434204102,36.4247398376465,36.4176254272461,36.4263305664062,36.4228515625,36.43115234375,36.4299201965332,36.4103355407715,36.3778343200684,36.3666801452637,36.3718338012695,36.3913764953613,36.4037971496582,36.3727531433105,36.3544273376465,36.3246231079102,36.2338829040527,36.0524063110352,36.0533485412598,36.0349349975586]}]],[[{"lng":[-81.8062210083008,-81.8171539306641,-81.8223114013672,-81.8502426147461,-81.8851928710938,-81.898567199707,-81.9065628051758,-81.8939971923828,-81.9111557006836,-81.8305740356445,-81.7304916381836,-81.7094573974609,-81.7403793334961,-81.7410736083984,-81.6330642700195,-81.5619812011719,-81.5408401489258,-81.4727554321289,-81.4508056640625,-81.4993209838867,-81.5461044311523,-81.6590042114258,-81.8062210083008],"lat":[36.1045608520508,36.1093864440918,36.1578559875488,36.1814727783203,36.1886367797852,36.1988563537598,36.221866607666,36.2669830322266,36.2907524108887,36.3346557617188,36.3293418884277,36.3337249755859,36.3618583679199,36.3917846679688,36.3406867980957,36.2735939025879,36.2725067138672,36.2343559265137,36.1906204223633,36.1360359191895,36.1113929748535,36.1175918579102,36.1045608520508]}]],[[{"lng":[-76.4805297851562,-76.5369567871094,-76.5755996704102,-76.5944900512695,-76.5710830688477,-76.5625076293945,-76.4603500366211,-76.3724212646484,-76.3242340087891,-76.2989273071289,-76.275505065918,-76.4805297851562],"lat":[36.0797920227051,36.0879249572754,36.1026573181152,36.239818572998,36.2777290344238,36.3405685424805,36.3738975524902,36.2523460388184,36.2336235046387,36.2142295837402,36.1103706359863,36.0797920227051]}]],[[{"lng":[-76.6887359619141,-76.6482162475586,-76.6042404174805,-76.5625076293945,-76.5710830688477,-76.5944900512695,-76.5755996704102,-76.5369567871094,-76.4805297851562,-76.4204254150391,-76.5230102539062,-76.5940017700195,-76.6490173339844,-76.6332092285156,-76.6901550292969,-76.7265090942383,-76.6887359619141],"lat":[36.2945175170898,36.315315246582,36.3149833679199,36.3405685424805,36.2777290344238,36.239818572998,36.1026573181152,36.0879249572754,36.0797920227051,36.0586051940918,36.0071678161621,36.0101318359375,36.0657081604004,36.0371170043945,36.0496101379395,36.1568222045898,36.2945175170898]}]],[[{"lng":[-81.9413452148438,-81.9613952636719,-81.944953918457,-81.947021484375,-81.9883651733398,-82.0050659179688,-82.0491027832031,-82.042724609375,-82.0623245239258,-82.0777587890625,-82.0204544067383,-81.9331130981445,-81.9111557006836,-81.8939971923828,-81.9065628051758,-81.898567199707,-81.8851928710938,-81.8502426147461,-81.8223114013672,-81.8171539306641,-81.8062210083008,-81.7322692871094,-81.8027877807617,-81.8598709106445,-81.8808135986328,-81.9008483886719,-81.9221496582031,-81.9413452148438],"lat":[35.9549751281738,35.9392204284668,35.9186134338379,35.9104118347168,35.9056434631348,35.9144401550293,35.9672355651855,36.0050086975098,36.0355262756348,36.1001396179199,36.129711151123,36.2633209228516,36.2907524108887,36.2669830322266,36.221866607666,36.1988563537598,36.1886367797852,36.1814727783203,36.1578559875488,36.1093864440918,36.1045608520508,36.0584754943848,35.9603309631348,35.9703407287598,35.9895248413086,35.9946517944336,35.9825134277344,35.9549751281738]}]],[[{"lng":[-80.4955444335938,-80.6895751953125,-80.8774108886719,-80.874382019043,-80.7826995849609,-80.7662963867188,-80.7525634765625,-80.7343673706055,-80.7240371704102,-80.6966552734375,-80.6674423217773,-80.6243133544922,-80.5905990600586,-80.5541534423828,-80.5450134277344,-80.5368804931641,-80.4763870239258,-80.4563446044922,-80.4408111572266,-80.4448089599609,-80.4919815063477,-80.5049896240234,-80.5082473754883,-80.4955444335938],"lat":[36.0432662963867,36.0454788208008,36.0524063110352,36.2338829040527,36.2485771179199,36.2618370056152,36.2582969665527,36.2647590637207,36.258472442627,36.259090423584,36.2461013793945,36.2730979919434,36.2682762145996,36.2784309387207,36.2766571044922,36.2567367553711,36.2547264099121,36.2425575256348,36.2194862365723,36.1233062744141,36.1076965332031,36.0949401855469,36.0708847045898,36.0432662963867]}]],[[{"lng":[-78.2545471191406,-78.266845703125,-78.3084106445312,-78.3295440673828,-78.3601150512695,-78.3944778442383,-78.4311447143555,-78.5480270385742,-78.5458526611328,-78.4925231933594,-78.4169540405273,-78.3808517456055,-78.3460464477539,-78.3087615966797,-78.1347198486328,-78.1096267700195,-78.058349609375,-78.0062866210938,-78.1309127807617,-78.2545471191406],"lat":[35.8155250549316,35.8483772277832,35.8934478759766,35.8878479003906,35.9186744689941,35.932300567627,35.9727172851562,36.0141258239746,36.0680885314941,36.1735877990723,36.1621742248535,36.2247505187988,36.2251815795898,36.2600402832031,36.2365837097168,36.213508605957,36.2113227844238,36.1959457397461,36.0214691162109,35.8155250549316]}]],[[{"lng":[-80.0381011962891,-80.1211471557617,-80.2098770141602,-80.2120666503906,-80.3343048095703,-80.3302841186523,-80.380241394043,-80.4115905761719,-80.4179763793945,-80.449951171875,-80.4648666381836,-80.484130859375,-80.4955444335938,-80.5082473754883,-80.5049896240234,-80.4919815063477,-80.4448089599609,-80.4408111572266,-80.4563446044922,-80.453010559082,-80.0256729125977,-80.0381011962891],"lat":[36.0061836242676,36.0215530395508,36.021427154541,35.9901237487793,35.9925651550293,35.9812355041504,35.9679298400879,35.9846115112305,36.0086326599121,36.0298385620117,36.0501937866211,36.0383224487305,36.0432662963867,36.0708847045898,36.0949401855469,36.1076965332031,36.1233062744141,36.2194862365723,36.2425575256348,36.2570877075195,36.2502326965332,36.0061836242676]}]],[[{"lng":[-79.5378189086914,-80.0426025390625,-80.0381011962891,-80.0256729125977,-79.5305786132812,-79.5378189086914],"lat":[35.8909683227539,35.9168128967285,36.0061836242676,36.2502326965332,36.236156463623,35.8909683227539]}]],[[{"lng":[-79.2461929321289,-79.2379913330078,-79.5409851074219,-79.5378189086914,-79.5305786132812,-79.5305099487305,-79.2584991455078,-79.2597732543945,-79.2708206176758,-79.2461929321289],"lat":[35.8681526184082,35.8372459411621,35.8369903564453,35.8909683227539,36.236156463623,36.2461357116699,36.2356872558594,36.0478935241699,35.9046020507812,35.8681526184082]}]],[[{"lng":[-76.7830657958984,-76.8105316162109,-76.8348388671875,-76.8567276000977,-76.8784866333008,-76.8984832763672,-76.9048080444336,-76.9360961914062,-76.9907302856445,-77.0044250488281,-77.0483169555664,-77.0341796875,-77.040885925293,-77.0662994384766,-77.0900115966797,-77.1886749267578,-77.1958923339844,-77.1933975219727,-77.2155380249023,-77.2138061523438,-77.2767181396484,-77.3152542114258,-77.3322067260742,-77.3098754882812,-77.2410354614258,-77.2574996948242,-77.2559127807617,-77.2704086303711,-77.2935104370117,-77.2986145019531,-77.2346115112305,-77.2176666259766,-77.1300735473633,-76.9947509765625,-76.9806900024414,-76.7450637817383,-76.7606735229492,-76.6937637329102,-76.7411270141602,-76.6972198486328,-76.7083206176758,-76.7285995483398,-76.7417221069336,-76.7612609863281,-76.7830657958984],"lat":[35.8592338562012,35.8813095092773,35.8842277526855,35.8357810974121,35.8172912597656,35.8141822814941,35.8679122924805,35.8945999145508,35.8778381347656,35.8618392944336,35.8574905395508,35.9143333435059,35.9317512512207,35.9360084533691,35.9252433776855,35.9312477111816,35.9359550476074,35.988094329834,35.9890670776367,36.0053634643555,36.0276985168457,36.0480842590332,36.0679817199707,36.0874443054199,36.1013031005859,36.1184730529785,36.130687713623,36.1555252075195,36.1628608703613,36.2115287780762,36.214599609375,36.2409820556641,36.2334632873535,36.2355804443359,36.2302360534668,36.2339172363281,36.1445922851562,35.9929695129395,35.9366264343262,35.9415435791016,35.9198341369629,35.9108581542969,35.8830757141113,35.8645439147949,35.8592338562012]}]],[[{"lng":[-79.0181350708008,-79.0953674316406,-79.2461929321289,-79.2708206176758,-79.2597732543945,-79.2584991455078,-79.1592712402344,-78.9510803222656,-79.0181350708008],"lat":[35.8578643798828,35.8539428710938,35.8681526184082,35.9046020507812,36.0478935241699,36.2356872558594,36.2336692810059,36.2338371276855,35.8578643798828]}]],[[{"lng":[-79.0181350708008,-78.9510803222656,-78.8068008422852,-78.8103561401367,-78.8040542602539,-78.7884140014648,-78.7491226196289,-78.7533950805664,-78.7281036376953,-78.7161712646484,-78.7041549682617,-78.7073440551758,-78.7341995239258,-78.7635040283203,-78.8082962036133,-78.8279571533203,-78.9057159423828,-79.0181350708008],"lat":[35.8578643798828,36.2338371276855,36.231575012207,36.114574432373,36.080940246582,36.062183380127,36.063591003418,36.0114631652832,36.0270729064941,36.0264930725098,35.9973258972168,35.9769439697266,35.9345855712891,35.9144744873047,35.9199256896973,35.8602294921875,35.8605155944824,35.8578643798828]}]],[[{"lng":[-78.1869277954102,-78.2056198120117,-78.2115478515625,-78.2340469360352,-78.2545471191406,-78.1309127807617,-78.0062866210938,-77.9851150512695,-77.9733963012695,-77.9558639526367,-77.9392318725586,-77.9224014282227,-77.917839050293,-77.8868408203125,-77.8113021850586,-77.8013687133789,-77.7471694946289,-77.7201843261719,-77.7005081176758,-77.8301467895508,-77.8449249267578,-77.873046875,-78.1869277954102],"lat":[35.7251129150391,35.7253952026367,35.7381935119629,35.745792388916,35.8155250549316,36.0214691162109,36.1959457397461,36.1774444580078,36.1890411376953,36.1837501525879,36.1875495910645,36.1777305603027,36.1567802429199,36.1444396972656,36.1358184814453,36.1442565917969,36.1464233398438,36.1341133117676,36.1441688537598,35.8544960021973,35.8355751037598,35.8447074890137,35.7251129150391]}]],[[{"lng":[-82.1188507080078,-82.1466522216797,-82.1410140991211,-82.1495666503906,-82.1846084594727,-82.1957702636719,-82.1990509033203,-82.2321014404297,-82.2406158447266,-82.2588577270508,-82.3016967773438,-82.3096923828125,-82.3296813964844,-82.3402633666992,-82.3444976806641,-82.3851776123047,-82.4084243774414,-82.3738555908203,-82.3119277954102,-82.2623062133789,-82.2077331542969,-82.154052734375,-82.1180801391602,-82.0777587890625,-82.0623245239258,-82.042724609375,-82.0491027832031,-82.0050659179688,-81.9883651733398,-81.9833450317383,-81.9908981323242,-82.0346832275391,-82.0979919433594,-82.1188507080078],"lat":[35.818531036377,35.8184814453125,35.9002723693848,35.9214477539062,35.9348754882812,35.9473762512207,36.0017700195312,36.0038757324219,35.984203338623,35.9856758117676,36.0293273925781,36.011474609375,36.0142631530762,36.0253982543945,36.070240020752,36.0785064697266,36.0753173828125,36.0986976623535,36.1221504211426,36.1203765869141,36.1470146179199,36.1396217346191,36.0962562561035,36.1001396179199,36.0355262756348,36.0050086975098,35.9672355651855,35.9144401550293,35.9056434631348,35.8875770568848,35.8724746704102,35.8535804748535,35.8438568115234,35.818531036377]}]],[[{"lng":[-77.6712188720703,-77.7331466674805,-77.7574920654297,-77.7550048828125,-77.7671356201172,-77.8301467895508,-77.7005081176758,-77.6856231689453,-77.6423568725586,-77.6062545776367,-77.5478820800781,-77.5366897583008,-77.5358581542969,-77.5251312255859,-77.4639739990234,-77.4382171630859,-77.4257431030273,-77.4053115844727,-77.3423690795898,-77.3581008911133,-77.3968048095703,-77.4056930541992,-77.4185791015625,-77.4269943237305,-77.4547119140625,-77.4732360839844,-77.5055160522461,-77.6712188720703],"lat":[35.6702651977539,35.7395477294922,35.7981033325195,35.824836730957,35.8368644714355,35.8544960021973,36.1441688537598,36.1466026306152,36.1267013549805,36.0973930358887,36.088493347168,36.0823631286621,36.0555725097656,36.0353851318359,36.0263824462891,36.0140342712402,35.9960632324219,35.9947166442871,35.9080047607422,35.8153114318848,35.8284187316895,35.8172645568848,35.8220863342285,35.8081970214844,35.793815612793,35.7996597290039,35.7667541503906,35.6702651977539]}]],[[{"lng":[-81.3281326293945,-81.3766555786133,-81.4175720214844,-81.457145690918,-81.5602569580078,-81.5933685302734,-81.7018280029297,-81.7484436035156,-81.7734756469727,-81.7756881713867,-81.8027877807617,-81.7322692871094,-81.8062210083008,-81.6590042114258,-81.5461044311523,-81.5173110961914,-81.4544372558594,-81.4296340942383,-81.3981475830078,-81.3887100219727,-81.3473510742188,-81.3218688964844,-81.3292617797852,-81.3395690917969,-81.330680847168,-81.3372116088867,-81.3281326293945],"lat":[35.7950592041016,35.7500343322754,35.7559051513672,35.7799339294434,35.775447845459,35.8131256103516,35.8687973022461,35.9207572937012,35.9222030639648,35.9439468383789,35.9603309631348,36.0584754943848,36.1045608520508,36.1175918579102,36.1113929748535,36.0954322814941,36.0830574035645,36.0656623840332,36.0560569763184,36.0375671386719,36.0153579711914,35.9893264770508,35.9892463684082,35.9292411804199,35.8758010864258,35.8276290893555,35.7950592041016]}]],[[{"lng":[-82.2792053222656,-82.2815780639648,-82.3218460083008,-82.3418502807617,-82.3453979492188,-82.3752517700195,-82.4058151245117,-82.4414825439453,-82.4870910644531,-82.4839477539062,-82.5069351196289,-82.4751968383789,-82.4084243774414,-82.3851776123047,-82.3444976806641,-82.3402633666992,-82.3296813964844,-82.3096923828125,-82.3016967773438,-82.2588577270508,-82.2406158447266,-82.2321014404297,-82.1990509033203,-82.1957702636719,-82.1846084594727,-82.1495666503906,-82.1410140991211,-82.1466522216797,-82.1188507080078,-82.1545944213867,-82.1642913818359,-82.211784362793,-82.2792053222656],"lat":[35.6975555419922,35.7202033996582,35.7398490905762,35.7635078430176,35.8051948547363,35.8164024353027,35.8139724731445,35.8822288513184,35.9048843383789,35.9476089477539,35.972541809082,35.9931755065918,36.0753173828125,36.0785064697266,36.070240020752,36.0253982543945,36.0142631530762,36.011474609375,36.0293273925781,35.9856758117676,35.984203338623,36.0038757324219,36.0017700195312,35.9473762512207,35.9348754882812,35.9214477539062,35.9002723693848,35.8184814453125,35.818531036377,35.7983703613281,35.7605247497559,35.7169876098633,35.6975555419922]}]],[[{"lng":[-77.1784591674805,-77.2076644897461,-77.2608184814453,-77.2645034790039,-77.3581008911133,-77.3423690795898,-77.4053115844727,-77.3322067260742,-77.3152542114258,-77.2767181396484,-77.2138061523438,-77.2155380249023,-77.1933975219727,-77.1958923339844,-77.1886749267578,-77.0900115966797,-77.0662994384766,-77.040885925293,-77.0341796875,-77.0483169555664,-77.0044250488281,-76.9907302856445,-76.9360961914062,-76.9048080444336,-76.8984832763672,-76.8784866333008,-76.8567276000977,-76.8348388671875,-76.8105316162109,-76.7830657958984,-76.8066787719727,-76.8052749633789,-76.8259582519531,-76.8383102416992,-76.9799118041992,-77.1612854003906,-77.1784591674805],"lat":[35.7321929931641,35.755126953125,35.759090423584,35.7827796936035,35.8153114318848,35.9080047607422,35.9947166442871,36.0679817199707,36.0480842590332,36.0276985168457,36.0053634643555,35.9890670776367,35.988094329834,35.9359550476074,35.9312477111816,35.9252433776855,35.9360084533691,35.9317512512207,35.9143333435059,35.8574905395508,35.8618392944336,35.8778381347656,35.8945999145508,35.8679122924805,35.8141822814941,35.8172912597656,35.8357810974121,35.8842277526855,35.8813095092773,35.8592338562012,35.8108406066895,35.7903747558594,35.7568817138672,35.7054557800293,35.6501274108887,35.7367782592773,35.7321929931641]}]],[[{"lng":[-78.9210739135742,-78.9988098144531,-78.9388885498047,-78.9444351196289,-78.9057159423828,-78.8279571533203,-78.8082962036133,-78.7635040283203,-78.7341995239258,-78.7073440551758,-78.7041549682617,-78.7161712646484,-78.7281036376953,-78.7533950805664,-78.7491226196289,-78.6955718994141,-78.5480270385742,-78.4311447143555,-78.3944778442383,-78.3601150512695,-78.3295440673828,-78.3084106445312,-78.266845703125,-78.2545471191406,-78.4776382446289,-78.7032165527344,-78.9210739135742],"lat":[35.578857421875,35.6013221740723,35.7614440917969,35.7701148986816,35.8605155944824,35.8602294921875,35.9199256896973,35.9144744873047,35.9345855712891,35.9769439697266,35.9973258972168,36.0264930725098,36.0270729064941,36.0114631652832,36.063591003418,36.0666465759277,36.0141258239746,35.9727172851562,35.932300567627,35.9186744689941,35.8878479003906,35.8934478759766,35.8483772277832,35.8155250549316,35.695613861084,35.5199394226074,35.578857421875]}]],[[{"lng":[-82.8959732055664,-82.8562698364258,-82.8086700439453,-82.7764434814453,-82.7735977172852,-82.7632293701172,-82.6438903808594,-82.628044128418,-82.6044006347656,-82.5922317504883,-82.6058044433594,-82.5993041992188,-82.5541458129883,-82.5069351196289,-82.4839477539062,-82.4870910644531,-82.4414825439453,-82.4058151245117,-82.5005493164062,-82.7663040161133,-82.8056259155273,-82.8432693481445,-82.8811111450195,-82.9075393676758,-82.9521865844727,-82.9430465698242,-82.9627532958984,-82.9068222045898,-82.9140701293945,-82.8959732055664],"lat":[35.9483604431152,35.9474258422852,35.9208717346191,35.9565734863281,35.9875030517578,35.9995460510254,36.0517234802246,36.0543403625488,36.0429878234863,36.0224494934082,36.003547668457,35.9632987976074,35.9561080932617,35.972541809082,35.9476089477539,35.9048843383789,35.8822288513184,35.8139724731445,35.7961273193359,35.6940002441406,35.6849060058594,35.6917266845703,35.6735610961914,35.7278480529785,35.7389984130859,35.7664642333984,35.7918510437012,35.8722152709961,35.9278678894043,35.9483604431152]}]],[[{"lng":[-80.7265167236328,-80.7762298583984,-80.9553451538086,-80.9510726928711,-80.9614334106445,-80.9312744140625,-81.0035781860352,-81.0547790527344,-81.0722045898438,-81.10888671875,-81.0491027832031,-80.9953460693359,-81.0205688476562,-80.9889526367188,-80.8774108886719,-80.6895751953125,-80.7059707641602,-80.7661209106445,-80.7265167236328],"lat":[35.507568359375,35.5068092346191,35.5091209411621,35.5286674499512,35.5435638427734,35.6195907592773,35.6970558166504,35.7134017944336,35.7436485290527,35.771900177002,35.8359680175781,35.9770774841309,36.0349349975586,36.0533485412598,36.0524063110352,36.0454788208008,35.8516578674316,35.6820373535156,35.507568359375]}]],[[{"lng":[-80.4567718505859,-80.4886932373047,-80.526725769043,-80.5723266601562,-80.6075592041016,-80.6349411010742,-80.7059707641602,-80.6895751953125,-80.4955444335938,-80.484130859375,-80.4648666381836,-80.449951171875,-80.4179763793945,-80.4115905761719,-80.380241394043,-80.3630676269531,-80.3617095947266,-80.4059371948242,-80.3865051269531,-80.3864364624023,-80.3954849243164,-80.4148025512695,-80.4260711669922,-80.4465026855469,-80.4577941894531,-80.4794235229492,-80.4728851318359,-80.4506454467773,-80.4567718505859],"lat":[35.7457962036133,35.7756156921387,35.7818069458008,35.8138122558594,35.8222579956055,35.840259552002,35.8516578674316,36.0454788208008,36.0432662963867,36.0383224487305,36.0501937866211,36.0298385620117,36.0086326599121,35.9846115112305,35.9679298400879,35.9421234130859,35.8958511352539,35.8843688964844,35.858570098877,35.8454132080078,35.839485168457,35.8448638916016,35.8303070068359,35.8306846618652,35.8224754333496,35.8332786560059,35.7861175537109,35.7648773193359,35.7457962036133]}]],[[{"lng":[-81.10888671875,-81.1272811889648,-81.1414031982422,-81.3281326293945,-81.3372116088867,-81.330680847168,-81.3395690917969,-81.3292617797852,-81.3218688964844,-81.2360076904297,-81.1574630737305,-81.1240615844727,-81.0840835571289,-81.0205688476562,-80.9953460693359,-81.0491027832031,-81.10888671875],"lat":[35.771900177002,35.7889671325684,35.8233184814453,35.7950592041016,35.8276290893555,35.8758010864258,35.9292411804199,35.9892463684082,35.9893264770508,36.0238227844238,36.0209808349609,36.0312843322754,36.0207672119141,36.0349349975586,35.9770774841309,35.8359680175781,35.771900177002]}]],[[{"lng":[-80.0644073486328,-80.1819076538086,-80.2092132568359,-80.251220703125,-80.2857818603516,-80.3255920410156,-80.326286315918,-80.3772583007812,-80.4567718505859,-80.4506454467773,-80.4728851318359,-80.4794235229492,-80.4577941894531,-80.4465026855469,-80.4260711669922,-80.4148025512695,-80.3954849243164,-80.3864364624023,-80.3865051269531,-80.4059371948242,-80.3617095947266,-80.3630676269531,-80.380241394043,-80.3302841186523,-80.3343048095703,-80.2120666503906,-80.2098770141602,-80.1211471557617,-80.0381011962891,-80.0426025390625,-80.0644073486328],"lat":[35.5056991577148,35.5051193237305,35.5745124816895,35.6248016357422,35.6369743347168,35.6831550598145,35.7140121459961,35.7106857299805,35.7457962036133,35.7648773193359,35.7861175537109,35.8332786560059,35.8224754333496,35.8306846618652,35.8303070068359,35.8448638916016,35.839485168457,35.8454132080078,35.858570098877,35.8843688964844,35.8958511352539,35.9421234130859,35.9679298400879,35.9812355041504,35.9925651550293,35.9901237487793,36.021427154541,36.0215530395508,36.0061836242676,35.9168128967285,35.5056991577148]}]],[[{"lng":[-81.8162841796875,-81.8656234741211,-81.985595703125,-81.9752960205078,-81.9329833984375,-81.9062576293945,-81.9062271118164,-81.9413452148438,-81.9221496582031,-81.9008483886719,-81.8808135986328,-81.8598709106445,-81.8027877807617,-81.7756881713867,-81.7734756469727,-81.7484436035156,-81.7018280029297,-81.5933685302734,-81.5602569580078,-81.457145690918,-81.4175720214844,-81.3766555786133,-81.4057235717773,-81.4956970214844,-81.5236434936523,-81.5625381469727,-81.5881118774414,-81.6439056396484,-81.6843185424805,-81.695198059082,-81.7008209228516,-81.7496109008789,-81.7656097412109,-81.8162841796875],"lat":[35.5743789672852,35.7188529968262,35.7999534606934,35.8182792663574,35.8275985717773,35.8479957580566,35.8920135498047,35.9549751281738,35.9825134277344,35.9946517944336,35.9895248413086,35.9703407287598,35.9603309631348,35.9439468383789,35.9222030639648,35.9207572937012,35.8687973022461,35.8131256103516,35.775447845459,35.7799339294434,35.7559051513672,35.7500343322754,35.697509765625,35.6065483093262,35.5612602233887,35.5553092956543,35.5617752075195,35.5532989501953,35.5654373168945,35.5716361999512,35.5956115722656,35.6017112731934,35.5842247009277,35.5743789672852]}]],[[{"lng":[-76.4084320068359,-76.6338195800781,-76.8383102416992,-76.8259582519531,-76.8052749633789,-76.8066787719727,-76.7830657958984,-76.7612609863281,-76.7417221069336,-76.7285995483398,-76.7083206176758,-76.6972198486328,-76.4094696044922,-76.3714828491211,-76.3835601806641,-76.3581924438477,-76.4086990356445,-76.4084320068359],"lat":[35.6991157531738,35.7030029296875,35.7054557800293,35.7568817138672,35.7903747558594,35.8108406066895,35.8592338562012,35.8645439147949,35.8830757141113,35.9108581542969,35.9198341369629,35.9415435791016,35.977466583252,35.9323425292969,35.9004135131836,35.8605918884277,35.7894821166992,35.6991157531738]}]],[[{"lng":[-76.1673049926758,-76.2102355957031,-76.232795715332,-76.2976303100586,-76.2734451293945,-76.4084320068359,-76.4086990356445,-76.3581924438477,-76.3835601806641,-76.3714828491211,-76.2137680053711,-76.0896377563477,-76.0260467529297,-76.0759124755859,-76.0430679321289,-76.1673049926758],"lat":[35.6968421936035,35.6043891906738,35.5946655273438,35.6116943359375,35.6894989013672,35.6991157531738,35.7894821166992,35.8605918884277,35.9004135131836,35.9323425292969,35.9768753051758,35.9629135131836,35.9204254150391,35.7568016052246,35.6838493347168,35.6968421936035]}]],[[{"lng":[-81.8162841796875,-81.8318939208984,-81.8402709960938,-81.8621520996094,-81.9670486450195,-82.0016021728516,-82.0361862182617,-82.1167526245117,-82.1707077026367,-82.2680053710938,-82.2906723022461,-82.2653427124023,-82.2842407226562,-82.2891845703125,-82.2792053222656,-82.211784362793,-82.1642913818359,-82.1545944213867,-82.1188507080078,-82.0979919433594,-82.0346832275391,-81.9908981323242,-81.9833450317383,-81.9883651733398,-81.947021484375,-81.944953918457,-81.9613952636719,-81.9413452148438,-81.9062271118164,-81.9062576293945,-81.9329833984375,-81.9752960205078,-81.985595703125,-81.8656234741211,-81.8162841796875],"lat":[35.5743789672852,35.5650634765625,35.5372428894043,35.5305480957031,35.5215873718262,35.5459747314453,35.5285873413086,35.5190048217773,35.5284652709961,35.5688591003418,35.5893020629883,35.6124877929688,35.6389045715332,35.6714897155762,35.6975555419922,35.7169876098633,35.7605247497559,35.7983703613281,35.818531036377,35.8438568115234,35.8535804748535,35.8724746704102,35.8875770568848,35.9056434631348,35.9104118347168,35.9186134338379,35.9392204284668,35.9549751281738,35.8920135498047,35.8479957580566,35.8275985717773,35.8182792663574,35.7999534606934,35.7188529968262,35.5743789672852]}]],[[{"lng":[-79.7649917602539,-80.0644073486328,-80.0426025390625,-79.5378189086914,-79.5409851074219,-79.5553588867188,-79.7649917602539],"lat":[35.5059356689453,35.5056991577148,35.9168128967285,35.8909683227539,35.8369903564453,35.5130500793457,35.5059356689453]}]],[[{"lng":[-79.5553588867188,-79.5409851074219,-79.2379913330078,-79.2461929321289,-79.0953674316406,-79.0181350708008,-78.9057159423828,-78.9444351196289,-78.9388885498047,-78.9988098144531,-78.9210739135742,-78.9744873046875,-79.0384979248047,-79.0672912597656,-79.1326065063477,-79.1942443847656,-79.2097320556641,-79.2205047607422,-79.2271347045898,-79.2760925292969,-79.2861328125,-79.3053665161133,-79.3150100708008,-79.312858581543,-79.3315505981445,-79.3429489135742,-79.5553588867188],"lat":[35.5130500793457,35.8369903564453,35.8372459411621,35.8681526184082,35.8539428710938,35.8578643798828,35.8605155944824,35.7701148986816,35.7614440917969,35.6013221740723,35.578857421875,35.5171546936035,35.5494613647461,35.598705291748,35.6246147155762,35.5751304626465,35.5534515380859,35.5508003234863,35.5653648376465,35.5302772521973,35.5444068908691,35.5427055358887,35.5391311645508,35.5268669128418,35.5219802856445,35.5102424621582,35.5130500793457]}]],[[{"lng":[-78.0653305053711,-78.1249008178711,-78.1624450683594,-78.1618041992188,-78.1869277954102,-77.873046875,-77.8449249267578,-77.8301467895508,-77.7671356201172,-77.7550048828125,-77.7574920654297,-77.7331466674805,-77.6712188720703,-77.6983261108398,-77.8262939453125,-77.8271789550781,-78.0021362304688,-78.0604858398438,-78.0653305053711],"lat":[35.5820388793945,35.5975189208984,35.6829833984375,35.7092933654785,35.7251129150391,35.8447074890137,35.8355751037598,35.8544960021973,35.8368644714355,35.824836730957,35.7981033325195,35.7395477294922,35.6702651977539,35.6539993286133,35.5742568969727,35.5829010009766,35.5759963989258,35.594669342041,35.5820388793945]}]],[[{"lng":[-80.2982406616211,-80.7265167236328,-80.7661209106445,-80.7059707641602,-80.6349411010742,-80.6075592041016,-80.5723266601562,-80.526725769043,-80.4886932373047,-80.4567718505859,-80.3772583007812,-80.326286315918,-80.3255920410156,-80.2857818603516,-80.251220703125,-80.2092132568359,-80.1819076538086,-80.2982406616211],"lat":[35.4948997497559,35.507568359375,35.6820373535156,35.8516578674316,35.840259552002,35.8222579956055,35.8138122558594,35.7818069458008,35.7756156921387,35.7457962036133,35.7106857299805,35.7140121459961,35.6831550598145,35.6369743347168,35.6248016357422,35.5745124816895,35.5051193237305,35.4948997497559]}]],[[{"lng":[-77.4738845825195,-77.5045623779297,-77.5039367675781,-77.5210494995117,-77.5234298706055,-77.5495758056641,-77.5646438598633,-77.6341323852539,-77.6983261108398,-77.6712188720703,-77.5055160522461,-77.4732360839844,-77.4547119140625,-77.4269943237305,-77.4185791015625,-77.4056930541992,-77.3968048095703,-77.3581008911133,-77.2645034790039,-77.2608184814453,-77.2076644897461,-77.1784591674805,-77.1774215698242,-77.1922378540039,-77.1955184936523,-77.1754379272461,-77.1872100830078,-77.1746826171875,-77.1520538330078,-77.1483459472656,-77.1193923950195,-77.1037673950195,-77.1473999023438,-77.1722106933594,-77.1949615478516,-77.2112045288086,-77.2405471801758,-77.2443008422852,-77.2641906738281,-77.2938003540039,-77.3542175292969,-77.3861618041992,-77.4014892578125,-77.4439468383789,-77.4738845825195],"lat":[35.4215278625488,35.4848327636719,35.5038909912109,35.5165061950684,35.5301780700684,35.5257263183594,35.5319404602051,35.5878257751465,35.6539993286133,35.6702651977539,35.7667541503906,35.7996597290039,35.793815612793,35.8081970214844,35.8220863342285,35.8172645568848,35.8284187316895,35.8153114318848,35.7827796936035,35.759090423584,35.755126953125,35.7321929931641,35.7144622802734,35.7120933532715,35.6999130249023,35.6762847900391,35.664306640625,35.6354103088379,35.6198844909668,35.5980033874512,35.5854988098145,35.5501861572266,35.5475883483887,35.5191230773926,35.4229545593262,35.3956413269043,35.3799858093262,35.354190826416,35.3501129150391,35.373950958252,35.3235549926758,35.3292617797852,35.3427696228027,35.3545951843262,35.4215278625488]}]],[[{"lng":[-80.9614334106445,-81.5236434936523,-81.4956970214844,-81.4057235717773,-81.3766555786133,-81.3281326293945,-81.1414031982422,-81.1272811889648,-81.10888671875,-81.0722045898438,-81.0547790527344,-81.0035781860352,-80.9312744140625,-80.9614334106445],"lat":[35.5435638427734,35.5612602233887,35.6065483093262,35.697509765625,35.7500343322754,35.7950592041016,35.8233184814453,35.7889671325684,35.771900177002,35.7436485290527,35.7134017944336,35.6970558166504,35.6195907592773,35.5435638427734]}]],[[{"lng":[-82.2581024169922,-82.3228759765625,-82.370964050293,-82.3648376464844,-82.3736877441406,-82.409065246582,-82.4749221801758,-82.5195159912109,-82.5269012451172,-82.5511322021484,-82.5639343261719,-82.6186828613281,-82.6689834594727,-82.7141571044922,-82.7438888549805,-82.7808837890625,-82.7947235107422,-82.7706298828125,-82.7720336914062,-82.8133392333984,-82.8231048583984,-82.8448638916016,-82.8655242919922,-82.8811111450195,-82.8432693481445,-82.8056259155273,-82.7663040161133,-82.5005493164062,-82.4058151245117,-82.3752517700195,-82.3453979492188,-82.3418502807617,-82.3218460083008,-82.2815780639648,-82.2792053222656,-82.2891845703125,-82.2842407226562,-82.2653427124023,-82.2906723022461,-82.2680053710938,-82.1707077026367,-82.2229080200195,-82.2276000976562,-82.2401580810547,-82.2581024169922],"lat":[35.4637298583984,35.4951591491699,35.4787712097168,35.4634590148926,35.4573783874512,35.468921661377,35.4444046020508,35.4439163208008,35.4283218383789,35.4268951416016,35.4384231567383,35.4376754760742,35.4551658630371,35.439151763916,35.4180335998535,35.4416885375977,35.4649696350098,35.5327339172363,35.5712928771973,35.6175346374512,35.6213874816895,35.6131477355957,35.6358032226562,35.6735610961914,35.6917266845703,35.6849060058594,35.6940002441406,35.7961273193359,35.8139724731445,35.8164024353027,35.8051948547363,35.7635078430176,35.7398490905762,35.7202033996582,35.6975555419922,35.6714897155762,35.6389045715332,35.6124877929688,35.5893020629883,35.5688591003418,35.5284652709961,35.5156936645508,35.4829216003418,35.4681549072266,35.4637298583984]}]],[[{"lng":[-78.5387420654297,-78.5394668579102,-78.6008224487305,-78.623176574707,-78.6887435913086,-78.7089691162109,-78.7032165527344,-78.4776382446289,-78.2545471191406,-78.2340469360352,-78.2115478515625,-78.2056198120117,-78.1869277954102,-78.1618041992188,-78.1624450683594,-78.1249008178711,-78.0653305053711,-78.1459732055664,-78.1568908691406,-78.1774597167969,-78.2090377807617,-78.2372283935547,-78.2681655883789,-78.3101272583008,-78.4166793823242,-78.4936218261719,-78.5387420654297],"lat":[35.3151168823242,35.3364639282227,35.4030303955078,35.4464149475098,35.5075263977051,35.514102935791,35.5199394226074,35.695613861084,35.8155250549316,35.745792388916,35.7381935119629,35.7253952026367,35.7251129150391,35.7092933654785,35.6829833984375,35.5975189208984,35.5820388793945,35.4289932250977,35.3506317138672,35.339599609375,35.3391761779785,35.3146324157715,35.3173599243164,35.2802848815918,35.2490768432617,35.2596206665039,35.3151168823242]}]],[[{"lng":[-82.7438888549805,-82.8332366943359,-82.9191131591797,-82.9536209106445,-82.9853591918945,-83.0385971069336,-83.0460433959961,-83.0877151489258,-83.1282348632812,-83.1423721313477,-83.181396484375,-83.1570205688477,-83.1803741455078,-83.1781005859375,-83.1951293945312,-83.1842956542969,-83.2152557373047,-83.2591247558594,-83.253303527832,-83.2438507080078,-83.1853485107422,-83.1436614990234,-83.1181869506836,-83.0599594116211,-82.9870071411133,-82.9627532958984,-82.9430465698242,-82.9521865844727,-82.9075393676758,-82.8811111450195,-82.8655242919922,-82.8448638916016,-82.8231048583984,-82.8133392333984,-82.7720336914062,-82.7706298828125,-82.7947235107422,-82.7808837890625,-82.7438888549805],"lat":[35.4180335998535,35.3155708312988,35.2907676696777,35.3085289001465,35.3563232421875,35.3899307250977,35.4069938659668,35.4463272094727,35.456615447998,35.4843940734863,35.5124092102051,35.5516624450684,35.582820892334,35.623291015625,35.6378173828125,35.6630783081055,35.6853981018066,35.6910095214844,35.7007064819336,35.7182159423828,35.7288856506348,35.7626838684082,35.7638092041016,35.7825775146484,35.7739906311035,35.7918510437012,35.7664642333984,35.7389984130859,35.7278480529785,35.6735610961914,35.6358032226562,35.6131477355957,35.6213874816895,35.6175346374512,35.5712928771973,35.5327339172363,35.4649696350098,35.4416885375977,35.4180335998535]}]],[[{"lng":[-75.7831726074219,-75.7731552124023,-75.5449676513672,-75.7027359008789,-75.7408676147461,-75.7831726074219],"lat":[36.2251930236816,36.2292556762695,35.7883605957031,36.049861907959,36.0503234863281,36.2251930236816]}],[{"lng":[-75.8914947509766,-75.9080276489258,-76.0212097167969,-75.9878540039062,-75.8180541992188,-75.7489624023438,-75.7293701171875,-75.779052734375,-75.8914947509766],"lat":[35.6312675476074,35.6656379699707,35.6690940856934,35.892707824707,35.9235191345215,35.8693389892578,35.6651725769043,35.578685760498,35.6312675476074]}],[{"lng":[-75.4912185668945,-75.5336227416992,-75.4569778442383,-75.5262985229492,-75.7492904663086,-75.6915664672852,-75.521484375,-75.4754180908203,-75.4912185668945],"lat":[35.6704978942871,35.768856048584,35.6173973083496,35.2279167175293,35.189826965332,35.2349891662598,35.2813568115234,35.5644950866699,35.6704978942871]}]],[[{"lng":[-77.1037673950195,-77.1193923950195,-77.1483459472656,-77.1520538330078,-77.1746826171875,-77.1872100830078,-77.1754379272461,-77.1955184936523,-77.1922378540039,-77.1774215698242,-77.1784591674805,-77.1612854003906,-76.9799118041992,-76.8383102416992,-76.6338195800781,-76.6089172363281,-76.6079483032227,-76.5858764648438,-76.5395965576172,-76.5189437866211,-76.4925384521484,-76.6381988525391,-76.6287689208984,-76.7053756713867,-77.1037673950195],"lat":[35.5501861572266,35.5854988098145,35.5980033874512,35.6198844909668,35.6354103088379,35.664306640625,35.6762847900391,35.6999130249023,35.7120933532715,35.7144622802734,35.7321929931641,35.7367782592773,35.6501274108887,35.7054557800293,35.7030029296875,35.6641540527344,35.635066986084,35.6094551086426,35.5940361022949,35.5776443481445,35.5417861938477,35.520336151123,35.4378967285156,35.4119338989258,35.5501861572266]}],[{"lng":[-76.6145172119141,-76.6402130126953,-76.8505706787109,-76.8976058959961,-77.1949615478516,-77.1722106933594,-77.1473999023438,-77.1037673950195,-76.9831848144531,-76.6949005126953,-76.6145172119141],"lat":[35.2729187011719,35.237247467041,35.2171669006348,35.2515716552734,35.4229545593262,35.5191230773926,35.5475883483887,35.5501861572266,35.4365005493164,35.3504257202148,35.2729187011719]}]],[[{"lng":[-83.3318176269531,-83.4246826171875,-83.4732284545898,-83.6791915893555,-83.6900863647461,-83.6893844604492,-83.6342163085938,-83.5988845825195,-83.5803833007812,-83.5867156982422,-83.63232421875,-83.6536102294922,-83.7449569702148,-83.8549880981445,-83.8798904418945,-83.9300765991211,-83.954704284668,-83.9100112915039,-83.8812255859375,-83.8302001953125,-83.77587890625,-83.6728744506836,-83.6138610839844,-83.56103515625,-83.5057983398438,-83.4582901000977,-83.387092590332,-83.3430252075195,-83.2984161376953,-83.2591247558594,-83.2152557373047,-83.1842956542969,-83.1951293945312,-83.1781005859375,-83.1803741455078,-83.1570205688477,-83.181396484375,-83.2266616821289,-83.2473068237305,-83.3098602294922,-83.35302734375,-83.3647155761719,-83.3298110961914,-83.3318176269531],"lat":[35.3193435668945,35.3153190612793,35.2916450500488,35.2722206115723,35.2877807617188,35.3082427978516,35.3340339660645,35.3623657226562,35.4010848999023,35.4276885986328,35.4367408752441,35.4210968017578,35.4414024353027,35.4501266479492,35.462028503418,35.4490165710449,35.4554595947266,35.4764785766602,35.5105857849121,35.5190620422363,35.552604675293,35.5649719238281,35.5717391967773,35.55517578125,35.5595512390137,35.5972785949707,35.6252174377441,35.6532592773438,35.6563262939453,35.6910095214844,35.6853981018066,35.6630783081055,35.6378173828125,35.623291015625,35.582820892334,35.5516624450684,35.5124092102051,35.5134544372559,35.5069885253906,35.4634742736816,35.4558982849121,35.413330078125,35.3639106750488,35.3193435668945]}]],[[{"lng":[-77.8051834106445,-77.8041076660156,-77.8305816650391,-77.8262939453125,-77.6983261108398,-77.6341323852539,-77.5646438598633,-77.5495758056641,-77.5234298706055,-77.5210494995117,-77.5039367675781,-77.5045623779297,-77.4738845825195,-77.4946746826172,-77.5349731445312,-77.5371780395508,-77.558723449707,-77.6218032836914,-77.6660537719727,-77.6844711303711,-77.6984176635742,-77.7606964111328,-77.8051834106445],"lat":[35.3645896911621,35.401798248291,35.4236259460449,35.5742568969727,35.6539993286133,35.5878257751465,35.5319404602051,35.5257263183594,35.5301780700684,35.5165061950684,35.5038909912109,35.4848327636719,35.4215278625488,35.4074440002441,35.4178237915039,35.4019813537598,35.3810882568359,35.3664855957031,35.339672088623,35.3459396362305,35.3698196411133,35.3619384765625,35.3645896911621]}]],[[{"lng":[-79.1824417114258,-79.2643203735352,-79.2847290039062,-79.3031997680664,-79.3245849609375,-79.3310928344727,-79.3624877929688,-79.3436737060547,-79.3429489135742,-79.3315505981445,-79.312858581543,-79.3150100708008,-79.3053665161133,-79.2861328125,-79.2760925292969,-79.2271347045898,-79.2205047607422,-79.2097320556641,-79.1942443847656,-79.1326065063477,-79.0672912597656,-79.0384979248047,-78.9744873046875,-79.1824417114258],"lat":[35.3045387268066,35.3445701599121,35.3928070068359,35.408805847168,35.4148292541504,35.4448204040527,35.4704132080078,35.491641998291,35.5102424621582,35.5219802856445,35.5268669128418,35.5391311645508,35.5427055358887,35.5444068908691,35.5302772521973,35.5653648376465,35.5508003234863,35.5534515380859,35.5751304626465,35.6246147155762,35.598705291748,35.5494613647461,35.5171546936035,35.3045387268066]}]],[[{"lng":[-81.9714431762695,-81.9645156860352,-82.0515823364258,-82.0841522216797,-82.1221694946289,-82.1551284790039,-82.2582778930664,-82.2787246704102,-82.2746810913086,-82.2581024169922,-82.2401580810547,-82.2276000976562,-82.2229080200195,-82.1707077026367,-82.1167526245117,-82.0361862182617,-82.0016021728516,-81.9670486450195,-81.8621520996094,-81.8402709960938,-81.8318939208984,-81.8162841796875,-81.7656097412109,-81.7496109008789,-81.7008209228516,-81.695198059082,-81.6843185424805,-81.6976852416992,-81.7371673583984,-81.7594909667969,-81.7653579711914,-81.8705902099609,-81.9714431762695],"lat":[35.1882820129395,35.2474250793457,35.323127746582,35.33935546875,35.3886184692383,35.397102355957,35.3933563232422,35.4356422424316,35.4516067504883,35.4637298583984,35.4681549072266,35.4829216003418,35.5156936645508,35.5284652709961,35.5190048217773,35.5285873413086,35.5459747314453,35.5215873718262,35.5305480957031,35.5372428894043,35.5650634765625,35.5743789672852,35.5842247009277,35.6017112731934,35.5956115722656,35.5716361999512,35.5654373168945,35.3532638549805,35.2541732788086,35.2206993103027,35.1824722290039,35.1831169128418,35.1882820129395]}]],[[{"lng":[-78.1631927490234,-78.1651763916016,-78.2574005126953,-78.3101272583008,-78.2681655883789,-78.2372283935547,-78.2090377807617,-78.1774597167969,-78.1568908691406,-78.1459732055664,-78.0653305053711,-78.0604858398438,-78.0021362304688,-77.8271789550781,-77.8262939453125,-77.8305816650391,-77.8041076660156,-77.8051834106445,-77.8300628662109,-77.8365783691406,-77.8876724243164,-77.894157409668,-77.9139785766602,-77.944694519043,-77.9639282226562,-78.0021591186523,-78.036506652832,-78.087532043457,-78.1631927490234],"lat":[35.1822891235352,35.1932182312012,35.2209510803223,35.2802848815918,35.3173599243164,35.3146324157715,35.3391761779785,35.339599609375,35.3506317138672,35.4289932250977,35.5820388793945,35.594669342041,35.5759963989258,35.5829010009766,35.5742568969727,35.4236259460449,35.401798248291,35.3645896911621,35.3423500061035,35.1717414855957,35.1549644470215,35.1441802978516,35.1599731445312,35.1682319641113,35.1640243530273,35.1864776611328,35.1856918334961,35.1701850891113,35.1822891235352]}]],[[{"lng":[-78.6127395629883,-78.716064453125,-78.812385559082,-78.8745727539062,-78.8860168457031,-78.9125823974609,-79.0958938598633,-79.1468353271484,-79.1690979003906,-79.2166519165039,-79.1824417114258,-78.9744873046875,-78.9210739135742,-78.7032165527344,-78.7089691162109,-78.6887435913086,-78.623176574707,-78.6008224487305,-78.5394668579102,-78.5387420654297,-78.5804138183594,-78.6127395629883],"lat":[35.2438316345215,35.2599792480469,35.2587203979492,35.2425346374512,35.2299346923828,35.2224655151367,35.1899642944336,35.2130432128906,35.2359008789062,35.2652778625488,35.3045387268066,35.5171546936035,35.578857421875,35.5199394226074,35.514102935791,35.5075263977051,35.4464149475098,35.4030303955078,35.3364639282227,35.3151168823242,35.2865753173828,35.2438316345215]}]],[[{"lng":[-81.3228225708008,-81.362174987793,-81.7653579711914,-81.7594909667969,-81.7371673583984,-81.6976852416992,-81.6843185424805,-81.6439056396484,-81.5881118774414,-81.5625381469727,-81.5236434936523,-81.5069580078125,-81.5155029296875,-81.4450759887695,-81.4074172973633,-81.3532638549805,-81.3647994995117,-81.3523483276367,-81.3194274902344,-81.3114242553711,-81.3228225708008],"lat":[35.1637573242188,35.1628532409668,35.1824722290039,35.2206993103027,35.2541732788086,35.3532638549805,35.5654373168945,35.5532989501953,35.5617752075195,35.5553092956543,35.5612602233887,35.5464973449707,35.5214233398438,35.4133682250977,35.359806060791,35.3273010253906,35.3103713989258,35.2751045227051,35.260498046875,35.1879501342773,35.1637573242188]}]],[[{"lng":[-80.9567718505859,-81.4450759887695,-81.5155029296875,-81.5069580078125,-81.5236434936523,-80.9614334106445,-80.9510726928711,-80.9553451538086,-80.9421997070312,-80.9538650512695,-80.9567718505859],"lat":[35.3974533081055,35.4133682250977,35.5214233398438,35.5464973449707,35.5612602233887,35.5435638427734,35.5286674499512,35.5091209411621,35.4511299133301,35.4360580444336,35.3974533081055]}]],[[{"lng":[-83.1062850952148,-83.1614990234375,-83.1450576782227,-83.1482009887695,-83.1771469116211,-83.1934814453125,-83.2179336547852,-83.2200393676758,-83.232780456543,-83.2639389038086,-83.2849807739258,-83.3106384277344,-83.3318176269531,-83.3298110961914,-83.3647155761719,-83.35302734375,-83.3098602294922,-83.2473068237305,-83.2266616821289,-83.181396484375,-83.1423721313477,-83.1282348632812,-83.0877151489258,-83.0460433959961,-83.0385971069336,-82.9853591918945,-82.9536209106445,-82.9191131591797,-82.9268569946289,-82.9787139892578,-82.9905853271484,-82.9820785522461,-83.0520858764648,-83.0438537597656,-83.0072784423828,-83.1062850952148],"lat":[35.0002784729004,35.0592231750488,35.0832862854004,35.091381072998,35.1114959716797,35.139217376709,35.158992767334,35.222526550293,35.2308082580566,35.2217712402344,35.226634979248,35.2599868774414,35.3193435668945,35.3639106750488,35.413330078125,35.4558982849121,35.4634742736816,35.5069885253906,35.5134544372559,35.5124092102051,35.4843940734863,35.456615447998,35.4463272094727,35.4069938659668,35.3899307250977,35.3563232421875,35.3085289001465,35.2907676696777,35.2279052734375,35.1857299804688,35.1581764221191,35.1275062561035,35.0634765625,35.0400695800781,35.0242042541504,35.0002784729004]}]],[[{"lng":[-79.6074676513672,-79.6483764648438,-79.7082366943359,-79.7649917602539,-79.5553588867188,-79.3429489135742,-79.3436737060547,-79.3624877929688,-79.3310928344727,-79.3245849609375,-79.3031997680664,-79.2847290039062,-79.2643203735352,-79.1824417114258,-79.2166519165039,-79.1690979003906,-79.1468353271484,-79.0958938598633,-79.0971755981445,-79.1220016479492,-79.1472778320312,-79.2352905273438,-79.3500518798828,-79.4553604125977,-79.4950637817383,-79.5529022216797,-79.5741958618164,-79.5756301879883,-79.6074676513672],"lat":[35.1591873168945,35.1897315979004,35.2784156799316,35.5059356689453,35.5130500793457,35.5102424621582,35.491641998291,35.4704132080078,35.4448204040527,35.4148292541504,35.408805847168,35.3928070068359,35.3445701599121,35.3045387268066,35.2652778625488,35.2359008789062,35.2130432128906,35.1899642944336,35.1768074035645,35.1701850891113,35.1735496520996,35.2032089233398,35.140796661377,35.0373573303223,35.0625076293945,35.0641059875488,35.0732650756836,35.1191253662109,35.1591873168945]}]],[[{"lng":[-81.0493011474609,-81.0239639282227,-81.0072784423828,-81.0015182495117,-81.0140533447266,-80.9796447753906,-80.9262771606445,-80.9377593994141,-80.9829254150391,-80.9567718505859,-80.9538650512695,-80.9421997070312,-80.9553451538086,-80.7762298583984,-80.7753829956055,-80.7616882324219,-80.7454376220703,-80.7627410888672,-80.6899871826172,-80.6594467163086,-80.5396423339844,-80.7596817016602,-80.7972183227539,-80.8401641845703,-80.8947143554688,-80.9277954101562,-81.0398864746094,-81.0655517578125,-81.0284423828125,-81.0490417480469,-81.0493011474609],"lat":[35.1515312194824,35.1490325927734,35.1632499694824,35.195987701416,35.2499008178711,35.3332672119141,35.3486747741699,35.3649253845215,35.3690948486328,35.3974533081055,35.4360580444336,35.4511299133301,35.5091209411621,35.5068092346191,35.4795722961426,35.4651374816895,35.4198570251465,35.4006805419922,35.3407592773438,35.2646751403809,35.2058181762695,35.03662109375,35.0281982421875,35.0020179748535,35.0597343444824,35.1012496948242,35.0372009277344,35.0664825439453,35.1054077148438,35.1320114135742,35.1515312194824]}]],[[{"lng":[-80.5029449462891,-80.5396423339844,-80.6594467163086,-80.6899871826172,-80.7627410888672,-80.7454376220703,-80.7616882324219,-80.7753829956055,-80.7762298583984,-80.7265167236328,-80.2982406616211,-80.3784713745117,-80.4817199707031,-80.5029449462891],"lat":[35.1869125366211,35.2058181762695,35.2646751403809,35.3407592773438,35.4006805419922,35.4198570251465,35.4651374816895,35.4795722961426,35.5068092346191,35.507568359375,35.4948997497559,35.384838104248,35.2106094360352,35.1869125366211]}]],[[{"lng":[-80.0714111328125,-80.0590667724609,-80.0658416748047,-80.0923461914062,-80.0935134887695,-80.0507507324219,-80.051887512207,-80.1163101196289,-80.1620712280273,-80.1819076538086,-80.0644073486328,-79.7649917602539,-79.7082366943359,-79.6483764648438,-79.6074676513672,-79.6372985839844,-79.6979522705078,-79.7598419189453,-79.8363189697266,-79.9094619750977,-79.9758224487305,-80.0106811523438,-80.0651779174805,-80.0714111328125],"lat":[35.1507415771484,35.1770782470703,35.2079429626465,35.2542266845703,35.291446685791,35.3663711547852,35.3768081665039,35.4643592834473,35.4747428894043,35.5051193237305,35.5056991577148,35.5059356689453,35.2784156799316,35.1897315979004,35.1591873168945,35.1538352966309,35.1730728149414,35.1673126220703,35.1738014221191,35.1557197570801,35.1507568359375,35.1371421813965,35.1365814208984,35.1507415771484]}]],[[{"lng":[-80.0714111328125,-80.1102752685547,-80.1316146850586,-80.1546859741211,-80.1591415405273,-80.1681289672852,-80.1901016235352,-80.2198867797852,-80.2475891113281,-80.2610931396484,-80.2751235961914,-80.3256759643555,-80.3487243652344,-80.3729476928711,-80.3987503051758,-80.428581237793,-80.4532852172852,-80.4832305908203,-80.5029449462891,-80.4817199707031,-80.3784713745117,-80.2982406616211,-80.1819076538086,-80.1620712280273,-80.1163101196289,-80.051887512207,-80.0507507324219,-80.0935134887695,-80.0923461914062,-80.0658416748047,-80.0590667724609,-80.0714111328125],"lat":[35.1507415771484,35.1938362121582,35.1711158752441,35.176082611084,35.1547393798828,35.149730682373,35.1678581237793,35.1591835021973,35.2045249938965,35.2040405273438,35.1931114196777,35.1739158630371,35.1706733703613,35.177864074707,35.1646118164062,35.1685943603516,35.1607856750488,35.1829109191895,35.1869125366211,35.2106094360352,35.384838104248,35.4948997497559,35.5051193237305,35.4747428894043,35.4643592834473,35.3768081665039,35.3663711547852,35.291446685791,35.2542266845703,35.2079429626465,35.1770782470703,35.1507415771484]}]],[[{"lng":[-82.5700302124023,-82.589225769043,-82.5962371826172,-82.6106109619141,-82.6081008911133,-82.7438888549805,-82.7141571044922,-82.6689834594727,-82.6186828613281,-82.5639343261719,-82.5511322021484,-82.5269012451172,-82.5195159912109,-82.4749221801758,-82.409065246582,-82.3736877441406,-82.3648376464844,-82.370964050293,-82.3228759765625,-82.2581024169922,-82.2746810913086,-82.2787246704102,-82.2582778930664,-82.315559387207,-82.34130859375,-82.345085144043,-82.3580322265625,-82.3508605957031,-82.3601226806641,-82.3713760375977,-82.3896102905273,-82.4379196166992,-82.4667434692383,-82.5246353149414,-82.5700302124023],"lat":[35.1494903564453,35.1849479675293,35.2424659729004,35.267578125,35.2930641174316,35.4180335998535,35.439151763916,35.4551658630371,35.4376754760742,35.4384231567383,35.4268951416016,35.4283218383789,35.4439163208008,35.4444046020508,35.468921661377,35.4573783874512,35.4634590148926,35.4787712097168,35.4951591491699,35.4637298583984,35.4516067504883,35.4356422424316,35.3933563232422,35.3096160888672,35.2854957580566,35.2431907653809,35.2429313659668,35.1926727294922,35.1829490661621,35.1827239990234,35.2082405090332,35.1695594787598,35.1735000610352,35.1545600891113,35.1494903564453]}]],[[{"lng":[-83.6956329345703,-83.7535629272461,-83.791877746582,-83.8368606567383,-83.9216537475586,-83.9608993530273,-83.9922027587891,-84.0038909912109,-84.0308609008789,-84.0292053222656,-84.0063095092773,-84.0126495361328,-83.954704284668,-83.9300765991211,-83.8798904418945,-83.8549880981445,-83.7449569702148,-83.6536102294922,-83.63232421875,-83.5867156982422,-83.5803833007812,-83.5988845825195,-83.6342163085938,-83.6893844604492,-83.6900863647461,-83.6791915893555,-83.6956329345703],"lat":[35.2508087158203,35.2371673583984,35.2486610412598,35.2467498779297,35.2189674377441,35.2153816223145,35.2325019836426,35.2620887756348,35.2925224304199,35.3252906799316,35.372859954834,35.4076232910156,35.4554595947266,35.4490165710449,35.462028503418,35.4501266479492,35.4414024353027,35.4210968017578,35.4367408752441,35.4276885986328,35.4010848999023,35.3623657226562,35.3340339660645,35.3082427978516,35.2877807617188,35.2722206115723,35.2508087158203]}]],[[{"lng":[-77.8365783691406,-77.8300628662109,-77.8051834106445,-77.7606964111328,-77.6984176635742,-77.6844711303711,-77.6660537719727,-77.6218032836914,-77.558723449707,-77.5371780395508,-77.5349731445312,-77.4946746826172,-77.4738845825195,-77.4439468383789,-77.4014892578125,-77.3861618041992,-77.4131546020508,-77.4502105712891,-77.4485092163086,-77.4296493530273,-77.4740982055664,-77.5283126831055,-77.5107727050781,-77.5330581665039,-77.599494934082,-77.7319259643555,-77.7447891235352,-77.754997253418,-77.7516021728516,-77.7641372680664,-77.8365783691406],"lat":[35.1717414855957,35.3423500061035,35.3645896911621,35.3619384765625,35.3698196411133,35.3459396362305,35.339672088623,35.3664855957031,35.3810882568359,35.4019813537598,35.4178237915039,35.4074440002441,35.4215278625488,35.3545951843262,35.3427696228027,35.3292617797852,35.3312187194824,35.302059173584,35.2856750488281,35.259838104248,35.2271919250488,35.2396965026855,35.1557693481445,35.1448822021484,35.0708694458008,35.0008354187012,35.0024452209473,35.0180816650391,35.0956764221191,35.1286125183105,35.1717414855957]}]],[[{"lng":[-82.8876953125,-83.0072784423828,-83.0438537597656,-83.0520858764648,-82.9820785522461,-82.9905853271484,-82.9787139892578,-82.9268569946289,-82.9191131591797,-82.8332366943359,-82.7438888549805,-82.6081008911133,-82.6106109619141,-82.5962371826172,-82.589225769043,-82.5700302124023,-82.6544952392578,-82.6860504150391,-82.6880340576172,-82.6973571777344,-82.7713470458984,-82.8876953125],"lat":[35.0553703308105,35.0242042541504,35.0400695800781,35.0634765625,35.1275062561035,35.1581764221191,35.1857299804688,35.2279052734375,35.2907676696777,35.3155708312988,35.4180335998535,35.2930641174316,35.267578125,35.2424659729004,35.1849479675293,35.1494903564453,35.119457244873,35.1214637756348,35.0978012084961,35.0912322998047,35.0854225158691,35.0553703308105]}]],[[{"lng":[-81.3228225708008,-81.3114242553711,-81.3194274902344,-81.3523483276367,-81.3647994995117,-81.3532638549805,-81.4074172973633,-81.4450759887695,-80.9567718505859,-80.9829254150391,-80.9377593994141,-80.9262771606445,-80.9796447753906,-81.0140533447266,-81.0015182495117,-81.0072784423828,-81.0239639282227,-81.0493011474609,-81.3228225708008],"lat":[35.1637573242188,35.1879501342773,35.260498046875,35.2751045227051,35.3103713989258,35.3273010253906,35.359806060791,35.4133682250977,35.3974533081055,35.3690948486328,35.3649253845215,35.3486747741699,35.3332672119141,35.2499008178711,35.195987701416,35.1632499694824,35.1490325927734,35.1515312194824,35.1637573242188]}]],[[{"lng":[-82.2101745605469,-82.2783279418945,-82.3207702636719,-82.3508605957031,-82.3580322265625,-82.345085144043,-82.34130859375,-82.315559387207,-82.2582778930664,-82.1551284790039,-82.1221694946289,-82.0841522216797,-82.0515823364258,-81.9645156860352,-81.9714431762695,-82.2101745605469],"lat":[35.1931266784668,35.1950073242188,35.1841888427734,35.1926727294922,35.2429313659668,35.2431907653809,35.2854957580566,35.3096160888672,35.3933563232422,35.397102355957,35.3886184692383,35.33935546875,35.323127746582,35.2474250793457,35.1882820129395,35.1931266784668]}]],[[{"lng":[-83.1062850952148,-83.5130081176758,-83.5272216796875,-83.5532379150391,-83.5620422363281,-83.6435623168945,-83.6448745727539,-83.7178039550781,-83.7395248413086,-83.7130813598633,-83.7223281860352,-83.6966934204102,-83.6956329345703,-83.6791915893555,-83.4732284545898,-83.4246826171875,-83.3318176269531,-83.3106384277344,-83.2849807739258,-83.2639389038086,-83.232780456543,-83.2200393676758,-83.2179336547852,-83.1934814453125,-83.1771469116211,-83.1482009887695,-83.1450576782227,-83.1614990234375,-83.1062850952148],"lat":[35.0002784729004,34.9920234680176,35.0197372436523,35.0357627868652,35.0559425354004,35.127498626709,35.1433601379395,35.1388092041016,35.1458396911621,35.1807518005371,35.1972732543945,35.2144355773926,35.2508087158203,35.2722206115723,35.2916450500488,35.3153190612793,35.3193435668945,35.2599868774414,35.226634979248,35.2217712402344,35.2308082580566,35.222526550293,35.158992767334,35.139217376709,35.1114959716797,35.091381072998,35.0832862854004,35.0592231750488,35.0002784729004]}]],[[{"lng":[-78.11376953125,-78.1137390136719,-78.1567611694336,-78.2568054199219,-78.2746124267578,-78.3059005737305,-78.3198852539062,-78.309700012207,-78.3223419189453,-78.3388671875,-78.3638153076172,-78.3866729736328,-78.4010009765625,-78.4992904663086,-78.5419921875,-78.5934753417969,-78.6342544555664,-78.6664123535156,-78.6581573486328,-78.645637512207,-78.6334838867188,-78.6452255249023,-78.6327133178711,-78.6352844238281,-78.6127395629883,-78.5804138183594,-78.5387420654297,-78.4936218261719,-78.4166793823242,-78.3101272583008,-78.2574005126953,-78.1651763916016,-78.1631927490234,-78.170280456543,-78.1552429199219,-78.1429824829102,-78.1954498291016,-78.1560974121094,-78.1358184814453,-78.11376953125],"lat":[34.7209854125977,34.6991806030273,34.6771545410156,34.5537528991699,34.5812759399414,34.6053581237793,34.6300926208496,34.6358528137207,34.6455726623535,34.6826057434082,34.6920433044434,34.7218894958496,34.7593307495117,34.8551063537598,34.9078712463379,34.9434700012207,34.9843673706055,35.0714683532715,35.093620300293,35.1025581359863,35.1537246704102,35.1897277832031,35.1972999572754,35.2109527587891,35.2438316345215,35.2865753173828,35.3151168823242,35.2596206665039,35.2490768432617,35.2802848815918,35.2209510803223,35.1932182312012,35.1822891235352,35.097484588623,35.0681915283203,34.9013252258301,34.7436256408691,34.7284774780273,34.7113456726074,34.7209854125977]}]],[[{"lng":[-76.943244934082,-76.9442291259766,-76.9898910522461,-76.9723358154297,-76.9666061401367,-76.9474334716797,-76.8976058959961,-76.8505706787109,-76.6402130126953,-76.6145172119141,-76.5912094116211,-76.5409774780273,-76.5071258544922,-76.6400909423828,-76.605224609375,-76.6242218017578,-76.6780090332031,-76.8494644165039,-76.9414901733398,-76.943244934082],"lat":[35.0700340270996,35.099723815918,35.1540451049805,35.1567726135254,35.1879653930664,35.2169914245605,35.2515716552734,35.2171669006348,35.237247467041,35.2729187011719,35.3120803833008,35.3036880493164,35.2487678527832,35.1725196838379,35.1385612487793,35.0643501281738,35.0240859985352,34.9822196960449,35.0274276733398,35.0700340270996]}]],[[{"lng":[-84.2910385131836,-84.225944519043,-84.1797256469727,-84.1016464233398,-84.0427780151367,-84.0308609008789,-84.0038909912109,-83.9922027587891,-83.9608993530273,-83.9216537475586,-83.8368606567383,-83.791877746582,-83.7535629272461,-83.6956329345703,-83.6966934204102,-83.7223281860352,-83.7130813598633,-83.7395248413086,-83.7553939819336,-83.8538818359375,-83.9073181152344,-83.9528503417969,-83.9597702026367,-83.9885482788086,-84.1182403564453,-84.3238525390625,-84.2910385131836],"lat":[35.2105445861816,35.261604309082,35.2409896850586,35.2455520629883,35.272575378418,35.2925224304199,35.2620887756348,35.2325019836426,35.2153816223145,35.2189674377441,35.2467498779297,35.2486610412598,35.2371673583984,35.2508087158203,35.2144355773926,35.1972732543945,35.1807518005371,35.1458396911621,35.148063659668,35.1257934570312,35.0704154968262,35.0484428405762,35.0191230773926,34.9890670776367,34.988224029541,34.9890098571777,35.2105445861816]}]],[[{"lng":[-78.4992904663086,-78.5174407958984,-78.831657409668,-78.8962631225586,-78.9125518798828,-78.9286270141602,-78.948486328125,-79.0336074829102,-79.0943145751953,-79.1068420410156,-79.0971755981445,-79.0958938598633,-78.9125823974609,-78.8860168457031,-78.8745727539062,-78.812385559082,-78.716064453125,-78.6127395629883,-78.6352844238281,-78.6327133178711,-78.6452255249023,-78.6334838867188,-78.645637512207,-78.6581573486328,-78.6664123535156,-78.6342544555664,-78.5934753417969,-78.5419921875,-78.4992904663086],"lat":[34.8551063537598,34.8435287475586,34.8443565368652,34.8309020996094,34.8274230957031,34.8797988891602,34.9026832580566,34.9538230895996,35.0378570556641,35.1192245483398,35.1768074035645,35.1899642944336,35.2224655151367,35.2299346923828,35.2425346374512,35.2587203979492,35.2599792480469,35.2438316345215,35.2109527587891,35.1972999572754,35.1897277832031,35.1537246704102,35.1025581359863,35.093620300293,35.0714683532715,34.9843673706055,34.9434700012207,34.9078712463379,34.8551063537598]}]],[[{"lng":[-77.0490036010742,-77.0574645996094,-77.0731353759766,-77.1006927490234,-77.1598205566406,-77.1642608642578,-77.1928405761719,-77.180061340332,-77.2062683105469,-77.2229919433594,-77.2854232788086,-77.3351364135742,-77.3699722290039,-77.4996337890625,-77.6392517089844,-77.682991027832,-77.7319259643555,-77.599494934082,-77.5330581665039,-77.5107727050781,-77.5283126831055,-77.4740982055664,-77.4234085083008,-77.2055130004883,-77.1792449951172,-77.1404495239258,-77.1311187744141,-77.1105880737305,-77.1033706665039,-77.1095352172852,-77.0490036010742],"lat":[34.8311920166016,34.8150482177734,34.8154411315918,34.7974967956543,34.7884979248047,34.7745170593262,34.8042869567871,34.8321495056152,34.8355026245117,34.8845100402832,34.9264030456543,34.9325408935547,34.914249420166,34.9625053405762,34.9840202331543,34.966724395752,35.0008354187012,35.0708694458008,35.1448822021484,35.1557693481445,35.2396965026855,35.2271919250488,35.1943054199219,35.0880546569824,35.0533599853516,35.0542373657227,35.0692825317383,35.0660362243652,35.0242614746094,34.8304290771484,34.8311920166016]}]],[[{"lng":[-80.8401641845703,-80.7972183227539,-80.7596817016602,-80.5396423339844,-80.5029449462891,-80.4832305908203,-80.4532852172852,-80.428581237793,-80.3987503051758,-80.3729476928711,-80.3487243652344,-80.3256759643555,-80.2751235961914,-80.3252792358398,-80.5615463256836,-80.8000640869141,-80.7856521606445,-80.8401641845703],"lat":[35.0020179748535,35.0281982421875,35.03662109375,35.2058181762695,35.1869125366211,35.1829109191895,35.1607856750488,35.1685943603516,35.1646118164062,35.177864074707,35.1706733703613,35.1739158630371,35.1931114196777,34.8147621154785,34.8152275085449,34.8161125183105,34.9406394958496,35.0020179748535]}]],[[{"lng":[-79.9199523925781,-80.3252792358398,-80.2751235961914,-80.2610931396484,-80.2475891113281,-80.2198867797852,-80.1901016235352,-80.1681289672852,-80.1591415405273,-80.1546859741211,-80.1316146850586,-80.1102752685547,-80.0714111328125,-80.0651779174805,-80.0584487915039,-80.0028076171875,-79.9752807617188,-79.9449157714844,-79.9274978637695,-79.9151382446289,-79.9084320068359,-79.882080078125,-79.8939208984375,-79.8670501708984,-79.8537063598633,-79.9014205932617,-79.9199523925781],"lat":[34.807918548584,34.8147621154785,35.1931114196777,35.2040405273438,35.2045249938965,35.1591835021973,35.1678581237793,35.149730682373,35.1547393798828,35.176082611084,35.1711158752441,35.1938362121582,35.1507415771484,35.1365814208984,35.1039848327637,35.0899276733398,35.0967330932617,35.1153373718262,35.115779876709,35.1066932678223,35.0789947509766,35.0444641113281,35.0072441101074,34.967716217041,34.9045829772949,34.852409362793,34.807918548584]}]],[[{"lng":[-79.3403015136719,-79.3606796264648,-79.3984146118164,-79.4553604125977,-79.3500518798828,-79.2352905273438,-79.1472778320312,-79.1220016479492,-79.0971755981445,-79.1068420410156,-79.0943145751953,-79.0336074829102,-79.1818542480469,-79.3403015136719],"lat":[34.8356094360352,34.944709777832,34.9962158203125,35.0373573303223,35.140796661377,35.2032089233398,35.1735496520996,35.1701850891113,35.1768074035645,35.1192245483398,35.0378570556641,34.9538230895996,34.8318824768066,34.8356094360352]}]],[[{"lng":[-76.5189437866211,-76.5395965576172,-76.5858764648438,-76.6079483032227,-76.6089172363281,-76.6338195800781,-76.4084320068359,-76.2734451293945,-76.2976303100586,-76.232795715332,-76.2102355957031,-76.1673049926758,-76.1047439575195,-76.0212097167969,-75.9080276489258,-75.8914947509766,-76.052978515625,-76.1816177368164,-76.4988021850586,-76.5337524414062,-76.4584350585938,-76.4465255737305,-76.5189437866211],"lat":[35.5776443481445,35.5940361022949,35.6094551086426,35.635066986084,35.6641540527344,35.7030029296875,35.6991157531738,35.6894989013672,35.6116943359375,35.5946655273438,35.6043891906738,35.6968421936035,35.6636390686035,35.6690940856934,35.6656379699707,35.6312675476074,35.4146041870117,35.3415336608887,35.4162178039551,35.4501991271973,35.5044250488281,35.5508651733398,35.5776443481445]}],[{"lng":[-76.0170059204102,-75.9756927490234,-75.8539352416992,-75.7637023925781,-75.8115386962891,-75.9039459228516,-76.0028762817383,-76.0170059204102],"lat":[35.0694351196289,35.1161842346191,35.1672821044922,35.1922836303711,35.1640205383301,35.1324920654297,35.0694351196289,35.0694351196289]}]],[[{"lng":[-77.6898345947266,-77.9266738891602,-77.9393081665039,-77.9891510009766,-78.0173645019531,-78.11376953125,-78.1358184814453,-78.1560974121094,-78.1954498291016,-78.1429824829102,-78.1552429199219,-78.170280456543,-78.1631927490234,-78.087532043457,-78.036506652832,-78.0021591186523,-77.9639282226562,-77.944694519043,-77.9139785766602,-77.894157409668,-77.8876724243164,-77.8365783691406,-77.7641372680664,-77.7516021728516,-77.754997253418,-77.7447891235352,-77.7319259643555,-77.682991027832,-77.6552658081055,-77.6898345947266],"lat":[34.7201957702637,34.7110061645508,34.7194061279297,34.7170944213867,34.728931427002,34.7209854125977,34.7113456726074,34.7284774780273,34.7436256408691,34.9013252258301,35.0681915283203,35.097484588623,35.1822891235352,35.1701850891113,35.1856918334961,35.1864776611328,35.1640243530273,35.1682319641113,35.1599731445312,35.1441802978516,35.1549644470215,35.1717414855957,35.1286125183105,35.0956764221191,35.0180816650391,35.0024452209473,35.0008354187012,34.966724395752,34.9189338684082,34.7201957702637]}]],[[{"lng":[-79.6859588623047,-79.9199523925781,-79.9014205932617,-79.8537063598633,-79.8670501708984,-79.8939208984375,-79.882080078125,-79.9084320068359,-79.9151382446289,-79.9274978637695,-79.9449157714844,-79.9752807617188,-80.0028076171875,-80.0584487915039,-80.0651779174805,-80.0106811523438,-79.9758224487305,-79.9094619750977,-79.8363189697266,-79.7598419189453,-79.6979522705078,-79.6372985839844,-79.6074676513672,-79.5756301879883,-79.5741958618164,-79.5529022216797,-79.4950637817383,-79.4553604125977,-79.5471038818359,-79.5730438232422,-79.5805206298828,-79.5655899047852,-79.6219253540039,-79.611442565918,-79.6601486206055,-79.6859588623047],"lat":[34.8052558898926,34.807918548584,34.852409362793,34.9045829772949,34.967716217041,35.0072441101074,35.0444641113281,35.0789947509766,35.1066932678223,35.115779876709,35.1153373718262,35.0967330932617,35.0899276733398,35.1039848327637,35.1365814208984,35.1371421813965,35.1507568359375,35.1557197570801,35.1738014221191,35.1673126220703,35.1730728149414,35.1538352966309,35.1591873168945,35.1191253662109,35.0732650756836,35.0641059875488,35.0625076293945,35.0373573303223,35.0000610351562,34.9783668518066,34.9447898864746,34.9075012207031,34.8495712280273,34.8177490234375,34.8178977966309,34.8052558898926]}]],[[{"lng":[-83.9379959106445,-83.9885482788086,-83.9597702026367,-83.9528503417969,-83.9073181152344,-83.8538818359375,-83.7553939819336,-83.7395248413086,-83.7178039550781,-83.6448745727539,-83.6435623168945,-83.5620422363281,-83.5532379150391,-83.5272216796875,-83.5130081176758,-83.5494155883789,-83.9379959106445],"lat":[34.9893913269043,34.9890670776367,35.0191230773926,35.0484428405762,35.0704154968262,35.1257934570312,35.148063659668,35.1458396911621,35.1388092041016,35.1433601379395,35.127498626709,35.0559425354004,35.0357627868652,35.0197372436523,34.9920234680176,34.9895362854004,34.9893913269043]}]],[[{"lng":[-76.8976058959961,-76.9474334716797,-76.9666061401367,-76.9723358154297,-76.9898910522461,-76.9442291259766,-76.943244934082,-77.0478820800781,-77.1046752929688,-77.1105880737305,-77.1311187744141,-77.1404495239258,-77.1792449951172,-77.2055130004883,-77.4234085083008,-77.4740982055664,-77.4296493530273,-77.4485092163086,-77.4502105712891,-77.4131546020508,-77.3861618041992,-77.3542175292969,-77.2938003540039,-77.2641906738281,-77.2443008422852,-77.2405471801758,-77.2112045288086,-77.1949615478516,-76.8976058959961],"lat":[35.2515716552734,35.2169914245605,35.1879653930664,35.1567726135254,35.1540451049805,35.099723815918,35.0700340270996,35.092155456543,35.0879859924316,35.0660362243652,35.0692825317383,35.0542373657227,35.0533599853516,35.0880546569824,35.1943054199219,35.2271919250488,35.259838104248,35.2856750488281,35.302059173584,35.3312187194824,35.3292617797852,35.3235549926758,35.373950958252,35.3501129150391,35.354190826416,35.3799858093262,35.3956413269043,35.4229545593262,35.2515716552734]}],[{"lng":[-76.6470489501953,-76.625617980957,-76.7502136230469,-77.0490036010742,-77.1095352172852,-77.1033706665039,-77.1105880737305,-77.0011901855469,-76.9131088256836,-76.6470489501953],"lat":[34.9063339233398,34.8906478881836,34.848274230957,34.8311920166016,34.8304290771484,35.0242614746094,35.0660362243652,35.0528869628906,34.9364700317383,34.9063339233398]}]],[[{"lng":[-79.4559707641602,-79.6675033569336,-79.6859588623047,-79.6601486206055,-79.611442565918,-79.6219253540039,-79.5655899047852,-79.5805206298828,-79.5730438232422,-79.5471038818359,-79.4553604125977,-79.3984146118164,-79.3606796264648,-79.3403015136719,-79.3444519042969,-79.3279724121094,-79.4559707641602],"lat":[34.634090423584,34.8006629943848,34.8052558898926,34.8178977966309,34.8177490234375,34.8495712280273,34.9075012207031,34.9447898864746,34.9783668518066,35.0000610351562,35.0373573303223,34.9962158203125,34.944709777832,34.8356094360352,34.808837890625,34.7724113464355,34.634090423584]}]],[[{"lng":[-77.5386352539062,-77.5763320922852,-77.6898345947266,-77.6552658081055,-77.682991027832,-77.6392517089844,-77.4996337890625,-77.3699722290039,-77.3351364135742,-77.2854232788086,-77.2229919433594,-77.2062683105469,-77.180061340332,-77.1928405761719,-77.1642608642578,-77.1489562988281,-77.12939453125,-77.1566009521484,-77.5386352539062],"lat":[34.4570083618164,34.4693450927734,34.7201957702637,34.9189338684082,34.966724395752,34.9840202331543,34.9625053405762,34.914249420166,34.9325408935547,34.9264030456543,34.8845100402832,34.8355026245117,34.8321495056152,34.8042869567871,34.7745170593262,34.764331817627,34.6849136352539,34.6606369018555,34.4570083618164]}]],[[{"lng":[-78.8645095825195,-78.9194717407227,-78.9507369995117,-78.9753646850586,-79.0022354125977,-79.0064163208008,-79.0294723510742,-79.0409469604492,-79.0745010375977,-79.4471435546875,-79.4559707641602,-79.3279724121094,-79.3444519042969,-79.3403015136719,-79.1818542480469,-79.0336074829102,-78.948486328125,-78.9286270141602,-78.9125518798828,-78.8962631225586,-78.8417053222656,-78.8033218383789,-78.8526000976562,-78.8456649780273,-78.8645095825195],"lat":[34.4771957397461,34.4536399841309,34.4493827819824,34.3991661071777,34.3880424499512,34.366268157959,34.3473739624023,34.3192977905273,34.304573059082,34.6190605163574,34.634090423584,34.7724113464355,34.808837890625,34.8356094360352,34.8318824768066,34.9538230895996,34.9026832580566,34.8797988891602,34.8274230957031,34.8309020996094,34.7350006103516,34.6851043701172,34.5665817260742,34.507453918457,34.4771957397461]}]],[[{"lng":[-77.1489562988281,-77.1642608642578,-77.1598205566406,-77.1006927490234,-77.0731353759766,-77.0574645996094,-77.0490036010742,-76.7502136230469,-76.625617980957,-76.6470489501953,-76.6700592041016,-76.5297775268555,-76.4450302124023,-76.4240570068359,-76.364013671875,-76.3294067382812,-76.3147125244141,-76.4699020385742,-76.6253128051758,-77.0505294799805,-77.1489562988281],"lat":[34.764331817627,34.7745170593262,34.7884979248047,34.7974967956543,34.8154411315918,34.8150482177734,34.8311920166016,34.848274230957,34.8906478881836,34.9063339233398,34.9699974060059,34.9746360778809,35.016658782959,34.9462471008301,34.9431114196777,34.9759559631348,34.9488143920898,34.7850608825684,34.7197532653809,34.698917388916,34.764331817627]}],[{"lng":[-76.5442733764648,-76.5551452636719,-76.5377502441406,-76.4844818115234,-76.4322967529297,-76.3746795654297,-76.2877044677734,-76.4692611694336,-76.5442733764648],"lat":[34.5878295898438,34.6106605529785,34.6139221191406,34.6976318359375,34.760684967041,34.8128700256348,34.877010345459,34.6932830810547,34.5878295898438]}]],[[{"lng":[-78.2614974975586,-78.3289794921875,-78.4379425048828,-78.6380844116211,-78.6598205566406,-78.6684951782227,-78.8118896484375,-78.8363571166992,-78.8645095825195,-78.8456649780273,-78.8526000976562,-78.8033218383789,-78.8417053222656,-78.8962631225586,-78.831657409668,-78.5174407958984,-78.4992904663086,-78.4010009765625,-78.3866729736328,-78.3638153076172,-78.3388671875,-78.3223419189453,-78.309700012207,-78.3198852539062,-78.3059005737305,-78.2746124267578,-78.2568054199219,-78.2624053955078,-78.2550582885742,-78.169677734375,-78.2614974975586],"lat":[34.3947868347168,34.3644218444824,34.3718414306641,34.4511451721191,34.4509429931641,34.4651298522949,34.4553184509277,34.4591979980469,34.4771957397461,34.507453918457,34.5665817260742,34.6851043701172,34.7350006103516,34.8309020996094,34.8443565368652,34.8435287475586,34.8551063537598,34.7593307495117,34.7218894958496,34.6920433044434,34.6826057434082,34.6455726623535,34.6358528137207,34.6300926208496,34.6053581237793,34.5812759399414,34.5537528991699,34.5279388427734,34.507381439209,34.4624519348145,34.3947868347168]}]],[[{"lng":[-78.0259246826172,-78.130241394043,-78.15478515625,-78.1846160888672,-78.1945648193359,-78.2614974975586,-78.169677734375,-78.2550582885742,-78.2624053955078,-78.2568054199219,-78.1567611694336,-78.1137390136719,-78.11376953125,-78.0173645019531,-77.9891510009766,-77.9393081665039,-77.9266738891602,-77.6898345947266,-77.5763320922852,-77.5386352539062,-77.5867614746094,-77.6097259521484,-77.6026153564453,-77.7505264282227,-77.8091430664062,-77.8283843994141,-77.888069152832,-77.9217834472656,-77.9439392089844,-77.9498138427734,-77.9790725708008,-77.9944534301758,-77.9866790771484,-78.0259246826172],"lat":[34.3287696838379,34.3641242980957,34.3622436523438,34.3481712341309,34.3510513305664,34.3947868347168,34.4624519348145,34.507381439209,34.5279388427734,34.5537528991699,34.6771545410156,34.6991806030273,34.7209854125977,34.728931427002,34.7170944213867,34.7194061279297,34.7110061645508,34.7201957702637,34.4693450927734,34.4570083618164,34.4209365844727,34.4350433349609,34.4126281738281,34.305046081543,34.359432220459,34.3879699707031,34.364070892334,34.3733139038086,34.3564376831055,34.3660850524902,34.3756866455078,34.3623161315918,34.339916229248,34.3287696838379]}]],[[{"lng":[-78.6557159423828,-79.0745010375977,-79.0409469604492,-79.0294723510742,-79.0064163208008,-79.0022354125977,-78.9753646850586,-78.9507369995117,-78.9194717407227,-78.8645095825195,-78.8363571166992,-78.8118896484375,-78.6684951782227,-78.6598205566406,-78.6380844116211,-78.4379425048828,-78.3289794921875,-78.2614974975586,-78.1945648193359,-78.1846160888672,-78.15478515625,-78.2610626220703,-78.3735733032227,-78.3611221313477,-78.4254302978516,-78.4927444458008,-78.5272369384766,-78.5442810058594,-78.5634307861328,-78.5877838134766,-78.6302719116211,-78.6347198486328,-78.6557159423828],"lat":[33.948673248291,34.304573059082,34.3192977905273,34.3473739624023,34.366268157959,34.3880424499512,34.3991661071777,34.4493827819824,34.4536399841309,34.4771957397461,34.4591979980469,34.4553184509277,34.4651298522949,34.4509429931641,34.4511451721191,34.3718414306641,34.3644218444824,34.3947868347168,34.3510513305664,34.3481712341309,34.3622436523438,34.2152633666992,34.2023506164551,34.1867218017578,34.1380653381348,34.158504486084,34.154857635498,34.134162902832,34.0589447021484,34.0306053161621,34.0102005004883,33.9779777526855,33.948673248291]}]],[[{"lng":[-77.9607315063477,-77.9658660888672,-77.9752807617188,-77.9831466674805,-78.0002212524414,-77.9953918457031,-78.0070190429688,-78.0113067626953,-78.0259246826172,-77.9866790771484,-77.9944534301758,-77.9790725708008,-77.9498138427734,-77.9439392089844,-77.9217834472656,-77.888069152832,-77.8283843994141,-77.8091430664062,-77.7505264282227,-77.864387512207,-77.894401550293,-77.9267578125,-77.9607315063477],"lat":[34.1892433166504,34.2422866821289,34.2433624267578,34.2616806030273,34.2678833007812,34.2827987670898,34.2848167419434,34.312614440918,34.3287696838379,34.339916229248,34.3623161315918,34.3756866455078,34.3660850524902,34.3564376831055,34.3733139038086,34.364070892334,34.3879699707031,34.359432220459,34.305046081543,34.1927375793457,34.0691795349121,34.0620346069336,34.1892433166504]}]],[[{"lng":[-78.6557159423828,-78.6347198486328,-78.6302719116211,-78.5877838134766,-78.5634307861328,-78.5442810058594,-78.5272369384766,-78.4927444458008,-78.4254302978516,-78.3611221313477,-78.3735733032227,-78.2610626220703,-78.15478515625,-78.130241394043,-78.0259246826172,-78.0113067626953,-78.0070190429688,-77.9953918457031,-78.0002212524414,-77.9831466674805,-77.9752807617188,-77.9658660888672,-77.9607315063477,-77.9585266113281,-78.0348052978516,-78.579719543457,-78.6557159423828],"lat":[33.948673248291,33.9779777526855,34.0102005004883,34.0306053161621,34.0589447021484,34.134162902832,34.154857635498,34.158504486084,34.1380653381348,34.1867218017578,34.2023506164551,34.2152633666992,34.3622436523438,34.3641242980957,34.3287696838379,34.312614440918,34.2848167419434,34.2827987670898,34.2678833007812,34.2616806030273,34.2433624267578,34.2422866821289,34.1892433166504,33.9925804138184,33.9142913818359,33.8819923400879,33.948673248291]}]]],null,null,{"interactive":true,"className":"","stroke":false,"color":["#EFF6FC","#F5FAFE","#DCE9F6","#F5F9FE","#ECF4FB","#ECF4FB","#F7FBFF","#F5FAFE","#F0F7FD","#EAF3FB","#F0F6FD","#D0E2F2","#EAF2FB","#EBF3FB","#E5EFF9","#D8E7F5","#EAF3FB","#DCEAF6","#EDF5FC","#F5FAFE","#F2F8FD","#F2F8FD","#EDF5FC","#ECF4FC","#5EA4D0","#2272B5","#CEE0F2","#EDF5FC","#DCE9F6","#A2CCE2","#D4E4F4","#F3F8FE","#D7E7F5","#D8E7F5","#F2F8FD","#EBF3FB","#3887C0","#F2F8FD","#D3E4F3","#EEF5FC","#EDF4FC","#C7DBEF","#D8E7F5","#F0F7FD","#F7FBFF","#E7F1FA","#D0E2F2","#EAF3FB","#D7E6F5","#CFE1F2","#CADEF0","#C3DAEE","#A9CFE5","#D4E4F4","#E6F0F9","#F4F9FE","#E0ECF8","#F3F8FE","#F1F7FD","#E4EFF9","#DDEBF7","#B6D4E9","#D6E6F4","#CCDFF1","#E5EFF9","#EFF6FC","#E1EDF8","#08306B","#D3E4F3","#EEF5FC","#E3EEF9","#E1EDF8","#F5FAFE","#D8E7F5","#EEF5FC","#90C2DE","#F4F9FE","#F2F8FD","#DDEAF7","#F4F9FE","#F0F6FD","#093F81","#F4F9FE","#D5E5F4","#EBF3FB","#EBF4FB","#F6FAFF","#E2EEF8","#E0ECF7","#F7FBFF","#C2D9ED","#E4EFF9","#68ABD5","#A4CCE3","#E3EEF9","#E9F2FA","#EEF5FC","#DAE8F6","#C7DBEF","#E5EFF9"],"weight":5,"opacity":0.5,"fill":true,"fillColor":["#EFF6FC","#F5FAFE","#DCE9F6","#F5F9FE","#ECF4FB","#ECF4FB","#F7FBFF","#F5FAFE","#F0F7FD","#EAF3FB","#F0F6FD","#D0E2F2","#EAF2FB","#EBF3FB","#E5EFF9","#D8E7F5","#EAF3FB","#DCEAF6","#EDF5FC","#F5FAFE","#F2F8FD","#F2F8FD","#EDF5FC","#ECF4FC","#5EA4D0","#2272B5","#CEE0F2","#EDF5FC","#DCE9F6","#A2CCE2","#D4E4F4","#F3F8FE","#D7E7F5","#D8E7F5","#F2F8FD","#EBF3FB","#3887C0","#F2F8FD","#D3E4F3","#EEF5FC","#EDF4FC","#C7DBEF","#D8E7F5","#F0F7FD","#F7FBFF","#E7F1FA","#D0E2F2","#EAF3FB","#D7E6F5","#CFE1F2","#CADEF0","#C3DAEE","#A9CFE5","#D4E4F4","#E6F0F9","#F4F9FE","#E0ECF8","#F3F8FE","#F1F7FD","#E4EFF9","#DDEBF7","#B6D4E9","#D6E6F4","#CCDFF1","#E5EFF9","#EFF6FC","#E1EDF8","#08306B","#D3E4F3","#EEF5FC","#E3EEF9","#E1EDF8","#F5FAFE","#D8E7F5","#EEF5FC","#90C2DE","#F4F9FE","#F2F8FD","#DDEAF7","#F4F9FE","#F0F6FD","#093F81","#F4F9FE","#D5E5F4","#EBF3FB","#EBF4FB","#F6FAFF","#E2EEF8","#E0ECF7","#F7FBFF","#C2D9ED","#E4EFF9","#68ABD5","#A4CCE3","#E3EEF9","#E9F2FA","#EEF5FC","#DAE8F6","#C7DBEF","#E5EFF9"],"fillOpacity":0.8,"smoothFactor":0.2,"noClip":false},null,null,null,{"interactive":false,"permanent":false,"direction":"auto","opacity":1,"offset":[0,0],"textsize":"10px","textOnly":false,"className":"","sticky":true},null]},{"method":"addLegend","args":[{"colors":["#F7FBFF , #CBDEF1 22.2680412371134%, #7EB8DA 45.6982193064667%, #3280BD 69.1283973758201%, #094388 92.5585754451734%, #08306B "],"labels":["5,000","10,000","15,000","20,000"],"na_color":null,"na_label":"NA","opacity":0.8,"position":"bottomright","type":"numeric","title":"BIR74","extra":{"p_1":0.222680412371134,"p_n":0.925585754451734},"layerId":null,"className":"info legend","group":null}]}],"limits":{"lat":[33.8819923400879,36.5896492004395],"lng":[-84.3238525390625,-75.4569778442383]}},"evals":[],"jsHooks":[]}</script> --- ## Exercise Use `ggplot` to create a choropleth map for the proportion of sudden infant deaths, for the period of July 1, 1974 to June 30, 1979. <img src="lec_11_files/figure-html/unnamed-chunk-38-1.png" width="100%" style="display: block; margin: auto;" /> ??? ```r nc %>% select(BIR74, SID74) %>% mutate(SID74_prop = SID74 / (BIR74 + SID74)) %>% ggplot() + geom_sf(aes(fill = SID74_prop)) + scale_fill_gradient(low = "#fff7f3", high = "#49006a") + labs(title = "July 1, 1974 to June 30, 1979", fill = "", subtitle = "Proportion of SID by county") + theme_void() + theme(plot.margin = margin(0, 1, 0, 1, "cm")) ``` --- class: inverse, center, middle # Map layers --- ## PMTW streams data The North Carolina Wildlife Resources Commission developed the Public Mountain Trout Waters (PMTW) digital data to enhance planning and management of trout waters. The GIS dataset depicts the trout regulations in effect on trout waters (streams and impoundments) managed under the PMTW program as listed in 2020-2021. We can directly download and unzip the shapefile via ```r download.file(str_c("https://opendata.arcgis.com/datasets/", "0cc135e6c6244c9e9646b45ee3cb6c1e_0.zip", "?outSR=%7B%22latestWkid%22%3A3857%2C%22", "wkid%22%3A102100%7D"), destfile = "data/pmtw.zip") unzip("data/pmtw.zip", exdir = "data/") ``` -- To see the available files ```r list.files(path = "data/", pattern = "PMTW_*") ``` ``` #> [1] "PMTW_Streams_2020.cpg" "PMTW_Streams_2020.dbf" "PMTW_Streams_2020.prj" #> [4] "PMTW_Streams_2020.shp" "PMTW_Streams_2020.shx" "PMTW_Streams_2020.xml" ``` --- ## Read in the shapefile ```r nc_pmtw <- st_read("data/PMTW_Streams_2020.shp", quiet = TRUE) ``` ```r glimpse(nc_pmtw) ``` ``` #> Rows: 1,858 #> Columns: 11 #> $ FID <int> 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17,… #> $ Excel_ID <int> 0, 0, 2, 4, 5, 6, 7, 8, 8, 9, 10, 11, 12, 12, 13, 20, 23, … #> $ Displ_Name <chr> "Big Sandy Creek", "Helton Creek", "Little River", "Cranbe… #> $ MHTW_Reach <chr> NA, NA, NA, NA, NA, NA, NA, NA, NA, NA, NA, NA, NA, NA, NA… #> $ FIRST_Reg_ <chr> "Helton Creek", "Helton Creek", "Little River", "Cranberry… #> $ FIRST_Reg1 <chr> "S.R. 1372 bridge to North Fork New River", "S.R. 1372 bri… #> $ FIRST_WRC_ <chr> "Delayed Harvest Trout Waters", "Delayed Harvest Trout Wat… #> $ FIRST_MHTW <chr> NA, NA, NA, NA, NA, NA, NA, NA, NA, NA, NA, NA, NA, NA, NA… #> $ WEB_Refere <chr> "http://www.ncwildlife.org/Fishing/LawsSafety/FishingRegul… #> $ Shape__Len <dbl> 37.44214, 17.63478, 4996.62215, 2925.42681, 5796.97361, 11… #> $ geometry <MULTILINESTRING [m]> MULTILINESTRING Z ((-901773..., MULTILINES… ``` --- ## Metadata for each `sf` object `nc`: ```r Simple feature collection with 100 features and 14 fields geometry type: MULTIPOLYGON dimension: XY bbox: xmin: -84.32385 ymin: 33.88199 xmax: -75.45698 ymax: 36.58965 *geographic CRS: NAD27 ``` `nc_pmtw`: ```r Simple feature collection with 1858 features and 10 fields geometry type: MULTILINESTRING dimension: XYZ bbox: xmin: -9383722 ymin: 4162088 xmax: -8947168 ymax: 4381832 z_range: zmin: 0 zmax: 0 *projected CRS: WGS 84 / Pseudo-Mercator ``` --- ## Check the CRS ```r st_crs(nc) ``` ```r Coordinate Reference System: User input: NAD27 wkt: GEOGCRS["NAD27", DATUM["North American Datum 1927", ELLIPSOID["Clarke 1866",6378206.4,294.978698213898, LENGTHUNIT["metre",1]]], PRIMEM["Greenwich",0, ANGLEUNIT["degree",0.0174532925199433]], CS[ellipsoidal,2], AXIS["latitude",north, ORDER[1], ANGLEUNIT["degree",0.0174532925199433]], AXIS["longitude",east, ORDER[2], ANGLEUNIT["degree",0.0174532925199433]], ID["EPSG",4267]] ``` --- .tiny[ ```r st_crs(nc_pmtw) ``` ```r Coordinate Reference System: User input: WGS 84 / Pseudo-Mercator wkt: PROJCRS["WGS 84 / Pseudo-Mercator", BASEGEOGCRS["WGS 84", DATUM["World Geodetic System 1984", ELLIPSOID["WGS 84",6378137,298.257223563, LENGTHUNIT["metre",1]]], PRIMEM["Greenwich",0, ANGLEUNIT["degree",0.0174532925199433]], ID["EPSG",4326]], CONVERSION["Popular Visualisation Pseudo-Mercator", METHOD["Popular Visualisation Pseudo Mercator", ID["EPSG",1024]], PARAMETER["Latitude of natural origin",0, ANGLEUNIT["degree",0.0174532925199433], ID["EPSG",8801]], PARAMETER["Longitude of natural origin",0, ANGLEUNIT["degree",0.0174532925199433], ID["EPSG",8802]], PARAMETER["False easting",0, LENGTHUNIT["metre",1], ID["EPSG",8806]], PARAMETER["False northing",0, LENGTHUNIT["metre",1], ID["EPSG",8807]]], CS[Cartesian,2], AXIS["easting (X)",east, ORDER[1], LENGTHUNIT["metre",1]], AXIS["northing (Y)",north, ORDER[2], LENGTHUNIT["metre",1]], USAGE[ SCOPE["unknown"], AREA["World - 85°S to 85°N"], BBOX[-85.06,-180,85.06,180]], ID["EPSG",3857]] ``` ] --- ## Coordinate reference systems (CRS) - CRS provide a standardized way of describing locations. - Different CRS arise from various ways data were gathered, the locations, and purposes of the data. - A CRS is comprised of - an ellipsoid, to define the earth's shape; - a datum, to define the origin and orientation of coordinate axes; - a projection, to go from 3D to 2D. - It is important that you transform your spatial data to a common CRS before plotting. --- ## Transform CRS ```r nc_pmtw <- st_transform(nc_pmtw, crs = st_crs(nc)) ``` -- Check they are equal: ```r st_crs(nc) == st_crs(nc_pmtw) ``` ``` #> [1] TRUE ``` --- class: center, middle <img src="lec_11_files/figure-html/unnamed-chunk-52-1.png" style="display: block; margin: auto;" /> --- ## Code: map overlay with `plot()` ```r plot(st_geometry(nc), axes = T, las = 1, main = "NC PMTW Streams", cex.main = 3, cex.lab = 2, cex.axis = 1.5) plot(st_geometry(nc_pmtw), * add = T, col = "#3BB3D0") legend("bottomleft", legend = "Trout streams", fill = "#3BB3D0") ``` --- class: center, middle <img src="lec_11_files/figure-html/unnamed-chunk-54-1.png" style="display: block; margin: auto;" /> --- ## Code: map overlay with `ggplot2` ```r # use some fancy fonts library(extrafont) loadfonts() ggplot(nc) + geom_sf(alpha = 0.2) + geom_sf(data = nc_pmtw, aes(color = FIRST_WRC_), size = 1.5) + labs(title = "NC PMTW Streams", color = "Stream designation", caption = str_c("Public mountain trout waters are only in ", "western North Carolina")) + theme_void() + xlim(c(-84.5, -80.5)) + # zoom long ylim(c(34.8, 36.8)) + # zoom lat # custom theme theme(legend.position = c(0.25, 0.80), plot.title = element_text(size = 50, family = "Impact"), legend.text = element_text(size = 16, family = "Comic Sans MS"), legend.title = element_text(size = 20, family = "Comic Sans MS"), plot.caption = element_text(size = 12, family = "Comic Sans MS")) ``` --- ## Exercise Recreate John Snow's cholera map showing the cholera death locations and the water pump locations. Download the data (you should have a directory named `data/`): ```r download.file("http://rtwilson.com/downloads/SnowGIS_SHP.zip", destfile = "data/john_snow.zip") unzip("data/john_snow.zip", exdir = "data/") ``` Read in the data: ```r cholera <- st_read("data/SnowGIS_SHP/Cholera_Deaths.shp", quiet = TRUE) pumps <- st_read("data/SnowGIS_SHP/Pumps.shp", quiet = TRUE) ``` Before you start, check that both `sf` objects have the same CRS. --- ## References 1. Interactive Viewing of Spatial Data in R. (2021). https://r-spatial.github.io/mapview/index.html. 2. "Leaflet For R - Introduction". Rstudio.Github.Io, 2021, https://rstudio.github.io/leaflet/. 3. Melanie Frazier. Coordinate Reference Systems in R. https://www.nceas.ucsb.edu/~frazier/RSpatialGuides/OverviewCoordinateReferenceSystems.pdf. 4. Simple Features for R. (2021). https://r-spatial.github.io/sf/.