class: center, middle, inverse, title-slide # Introduction & R Fundamentals ## Statistical Computing & Programming ### Shawn Santo ### 05-13-20 --- class: center, inverse, middle # Introduction --- ## Who am I? - Shawn Santo - [shawn.santo@duke.edu](mailto:shawn.santo@duke.edu) - Office hours (Zoom) - Wednesday 1:00 – 2:00pm, Zoom - Friday 8:30 – 9:30am, Zoom *All times listed are in Eastern Time.* --- ## Who else is involved? - [Pierre Gardan](https://stat.duke.edu/people/pierre-gardan-0) - [pierre.gardan@duke.edu](mailto:pierre.gardan@duke.edu) - Office hours (Zoom): Monday 10:00 - 11:00am, Thursday 3:30 - 4:30pm - [Abbey List]() - [abbey.list@duke.edu](mailto:abbey.list@duke.edu) - Office hours (Zoom): Tuesday 1:00 - 2:00pm, Thursday 1:00 - 2:00pm - [Bo Liu](https://stat.duke.edu/people/bo-liu-0) - [bo.liu1997@duke.edu](mailto:bo.liu1997@duke.edu) - Office hours (Zoom): Monday 4:30 - 5:30pm, Friday 3:00 - 4:00pm *All times listed are in Eastern Time.* --- ## What is statistical computing / programming? .middle.center[ <img src="images/statistical-computing-venn.png" height="400px"> ] *Source:* Deborah Nolan & Duncan Temple Lang (2010) Computing in the Statistics Curricula, The American Statistician, 64:2, 97-107, DOI: 10.1198/tast.2010.09132 --- ## What you will learn .pull-left[ - Fundamentals of R - S3 objects - Data visualization with package `ggplot2` - Package `tidyverse` - Web scraping - Web based applications with RShiny - Wrangling and managing big data - SQL and databases ] .pull-right[ - Data types and functions - Parallelization - Git and GitHub - Shell - Reproducible reports with R Markdown - Debugging and testing - Spark - Make ] <br><br> [Full course schedule](http://www2.stat.duke.edu/courses/Summer20/sta323.001-1/schedule.html) --- ## Why this class matters .middle.center[ 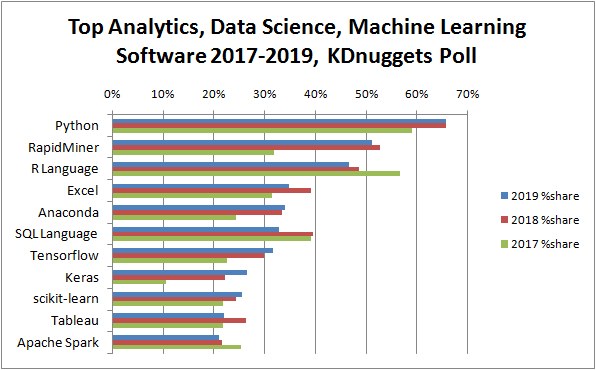 ] *Source:* https://www.kdnuggets.com/2019/05/poll-top-data-science-machine-learning-platforms.html --- ## Why this class matters ### Some 2020 internships: - <b>Mayo Clinic :</b> <i> Interns will work with statisticians, bioinformaticists, and clinical investigators on research projects in areas such as clinical trials, statistical genetics, and bioinformatics. Experience with SAS and/or R preferred.</i> - <b>Netflix (Science and Analytics):</b> <i> Comfortable coding in at least one language (e.g., R, Python, Java, Scala, C++), experience preferred with version control (e.g., git), great communication skills, both oral and written.</i> - <b>Two Sigma:</b> <i> Use the scientific method to develop sophisticated investment models and shape our insights into how the markets will behave. Create and test complex investment ideas and partner with our engineers to test your theories. You should possess the following qualifications: Demonstrate intermediate skills in at least one programming language, performed an in-depth research project, examining real-world data, are an independent thinker who can creatively approach data analysis and communicate complex ideas clearly.</i> *Source:* https://stattrak.amstat.org/2019/12/01/2020-internship-listings/ --- ## Why this class matters .middle.center[ <img src="images/top-10-ds-language.png" height="350px"> ] 100 data science job ads from LinkedIn across USA, UK, Canada, and Australia between April 22, 2019 and May 5, 2019 *Source*: https://towardsdatascience.com/which-programming-language-should-data-scientists-learn-first-aac4d3fd3038 --- ## Text toolkit .middle.center[  ] These are recommended textbooks - **all are available for free online**. There is no required textbook for this course. --- ## Software toolkit .middle.center[  ] --- ## Course structure This class is about you doing as opposed to you just watching or listening. Video lectures and labs will be interactive. My role as instructor is to introduce you to new tools and techniques, but it is up to you to take them and make use of them. If you only read the code and never run it or experiment with it, then you will not get much out of this course. Most slides will include supplemental resources for you to delve deeper in the topic of discussion. Occasionally, there will be readings assigned. To be successful in this course as an undergraduate student, you will need to commit up to 20 hours per week of your time. If you are a graduate student, you need to commit up to 25 hours per week of your time. In this online summer version, we will cover the same topics and at the same depth as what is covered in the typical 15-week semester. I have detailed a course schedule that you should follow in order to be successful in this course. --- ## Grading | Grade Item | Percentage | |-----------:|:----------:| | Homework | 40% | | Exam | 25% | | Project | 25% | | Labs | 10% | <br> <br> The exact ranges for letter grades may be curved and cutoffs will be determined at the end of the semester. However, if you have a cumulative numerical average of 90 - 100, you are guaranteed at least an A-, 80 - 89 at least a B-, 70 - 79 at least a C-, and so on. --- ## Teams I will construct teams based on the results of a survey you complete. Team expectations: - Each member must commit to giving equal effort. - Each member must read, run, and understand all code in a final submission. - Each member must honestly complete the intragroup peer evaluation. --- ## Policies - sharing / reusing code - Similar reproducible examples (reprex) exist online that will help you answer many of the questions posed on labs and homework assignments. Use of these resources is allowed unless it is written explicitly on the assignment. - You must always cite any code you copy or use as inspiration. Copied code without citation is plagiarism and will result in a 0 for the assignment. There may also be additional punitive measures taken depending on the severity of plagiarism. - Copying and citing a large amount of code to satisfy a main objective of an assignment will result in a 0 for the assignment. - Discussion (not code sharing / copying) with other students and groups is allowed unless it is written explicitly on the assignment. - Carefully read each assignment so you know what is permitted and what is not. If you are ever unsure what is allowed, please ask myself or one of the TAs. --- ## Getting help - Post your content and course related questions on Slack - Set up a Zoom meeting with myself or one of the TAs - Email me or one of the TAs --- ## Links to bookmark - Course page: http://www2.stat.duke.edu/courses/Summer20/sta323.001-1/ - GitHub organization: https://github.com/sta323-523-su20 - To access the DSS RStudio servers, use `AnyConnect` to connect with Duke's VPN. Navigate to: - Server http://pawn.stat.duke.edu:8787/ for undergraduate students - Server http://rook.stat.duke.edu:8787/ for graduate students --- ## To do list Before moving to the next section, please 1. create a GitHub account, - https://github.com/join 2. join Slack, 3. verify you can log-in to the Department's RStudio servers, 4. complete the first-day survey. Links for the necessary items were sent via email on 05-12-20. --- class: center, middle, inverse # Fundamentals of R --- ## Supplementary materials Companion videos - [Vectors](https://warpwire.duke.edu/w/YcADAA/) - [Operators, vectorization, and length coercion](https://warpwire.duke.edu/w/W8ADAA/) - [Control flow](https://warpwire.duke.edu/w/V8ADAA/) - [Error action](https://warpwire.duke.edu/w/WcADAA/) - [Loops](https://warpwire.duke.edu/w/Y8ADAA/) - [Introduction to using RMarkdown](https://warpwire.duke.edu/w/XcADAA/) Additional resources - [Google’s R Style Guide](https://google.github.io/styleguide/Rguide.html) - [Hadley's R Style Guide](http://r-pkgs.had.co.nz/style.html) - [Sections 3.1 – 3.2](https://adv-r.hadley.nz/vectors-chap.html) Advanced R - [Chapter 5](https://adv-r.hadley.nz/control-flow.html) Advanced R --- class: inverse, center, middle # Vectors --- ## Vectors The fundamental building block of data in R is a vector (collections of related values, objects, other data structures, etc). -- R has two types of vectors: * **atomic** vectors - homogeneous collections of the *same* type (e.g. all logical values, all numbers, or all character strings). * **generic** vectors - heterogeneous collections of *any* type of R object, even other lists (meaning they can have a hierarchical/tree-like structure). <br><br> I will use the term component or element when referring to a value inside a vector. --- ## Vector interrelationships .middle.center[ <img src="images/vector-types.png" height="400px"> ] *Source*: https://r4ds.had.co.nz/vectors.html --- ## Atomic vectors R has six atomic vector types: .center[ `logical`, `double`, `integer`, `character`, `complex`, `raw` ] In this course we will mostly work with the first four. You will rarely work with the last two types - complex and raw. ```r x <- c(T, F, TRUE, FALSE) typeof(x) ``` ``` #> [1] "logical" ``` ```r y <- c("a", "few", "more", "slides") typeof(y) ``` ``` #> [1] "character" ``` --- ## Coercion hierarchy If you try to combine components of different types into a single atomic vector, R will try to coerce all elements so they can be represented as the simplest type. .center[ `character` `\(\rightarrow\)` `double` `\(\rightarrow\)` `integer` `\(\rightarrow\)` `logical` ] ```r x <- c(T, 5, F, 0, 1) y <- c("a", 1, T) z <- c(3.0, 4L, 0L) ``` -- .pull-left[ ```r x ``` ``` #> [1] 1 5 0 0 1 ``` ```r y ``` ``` #> [1] "a" "1" "TRUE" ``` ```r z ``` ``` #> [1] 3 4 0 ``` ] .pull-right[ ```r typeof(x) ``` ``` #> [1] "double" ``` ```r typeof(y) ``` ``` #> [1] "character" ``` ```r typeof(z) ``` ``` #> [1] "double" ``` ] --- ## Concatenation One way to construct atomic vectors is with function `c()`. ```r c(1, 0, 1, 1, 6) ``` ``` #> [1] 1 0 1 1 6 ``` ```r c(c(3, 4), c(10, TRUE)) ``` ``` #> [1] 3 4 10 1 ``` ```r c(pi) ``` ``` #> [1] 3.141593 ``` --- class: inverse, center, middle # Operators, vectorization, and length coercion --- ## Logical (Boolean) operators | Operator | Operation | Vectorized? |:-----------------------------:|:-------------:|:------------: | <code>x | y</code> | or | Yes | `x & y` | and | Yes | `!x` | not | Yes | <code>x || y</code> | or | No | `x && y` | and | No |`xor(x,y)` | exclusive or | Yes <br><br> -- What do we mean if we say a function or operation is vectorized? --- ## Boolean examples ```r x <- c(T, F, T, T) y <- c(F, F, T, F) ``` .pull-left[ ```r !x ``` ``` #> [1] FALSE TRUE FALSE FALSE ``` ```r x | y ``` ``` #> [1] TRUE FALSE TRUE TRUE ``` ```r x || y ``` ``` #> [1] TRUE ``` ] .pull-right[ ```r x & y ``` ``` #> [1] FALSE FALSE TRUE FALSE ``` ```r x && y ``` ``` #> [1] FALSE ``` ```r xor(x, y) ``` ``` #> [1] TRUE FALSE FALSE TRUE ``` ] --- ## Comparison operators | Operator | Comparison | Vectorized? |:----------:|:--------------------------:|:----------------: | `x < y` | less than | Yes | `x > y` | greater than | Yes | `x <= y` | less than or equal to | Yes | `x >= y` | greater than or equal to | Yes | `x != y` | not equal to | Yes | `x == y` | equal to | Yes | `x %in% y` | contains | Yes (over `x`) --- ## Comparison examples ```r x <- c(4, 10, -5) y <- c(0, 51, 9 / 5) z <- c("four", "for", "4") ``` .pull-left[ ```r x > y ``` ``` #> [1] TRUE FALSE FALSE ``` ```r x != y ``` ``` #> [1] TRUE TRUE TRUE ``` ] .pull-right[ ```r x == z ``` ``` #> [1] FALSE FALSE FALSE ``` ```r x %in% z ``` ``` #> [1] TRUE FALSE FALSE ``` ] --- ## What else is vectorized? - Most of the mathematical operators - Many functions built-in to R and created by user's in packages ```r a <- c(0, -3, sqrt(75)) b <- c(1, 3, 2) ``` .pull-left[ ```r a + b ``` ``` #> [1] 1.00000 0.00000 10.66025 ``` ```r a ^ b ``` ``` #> [1] 0 -27 75 ``` ] .pull-right[ ```r rnorm(n = 3, mean = a, sd = b) ``` ``` #> [1] 0.1255788 -4.7301223 7.4654714 ``` ```r exp(a / b) ``` ``` #> [1] 1.0000000 0.3678794 75.9539335 ``` ] --- ## Length coercion (vector recycling) The shorter of two atomic vectors in an operation is recycled until it is the same length as the longer atomic vector. ```r x <- c(2, 4, 6) y <- c(1, 1, 1, 2, 2) ``` .small-text[ ```r x > y ``` ``` #> [1] TRUE TRUE TRUE FALSE TRUE ``` ```r x == y ``` ``` #> [1] FALSE FALSE FALSE TRUE FALSE ``` ```r 10 / x ``` ``` #> [1] 5.000000 2.500000 1.666667 ``` ] --- class: inverse, center, middle # Control flow --- ## Conditional control flow Conditional (choice) control flow is governed by `if` and `switch()`. .pull-left[ ```r if (condition) { # code to run # when condition is # TRUE } ``` ] .pull-right[ ```r if (TRUE) { print("The condition must have been true!") } ``` ] --- ## `if` examples ```r if (1 > 0) { print("Yes, 1 is greater than 0.") } ``` ``` #> [1] "Yes, 1 is greater than 0." ``` -- ```r x <- c(1, 2, 3, 4) if (3 %in% x) { print("Yes, 3 is in x.") } ``` ``` #> [1] "Yes, 3 is in x." ``` -- ```r if (-6) { print("Other types are coerced to logical if possible.") } ``` ``` #> [1] "Other types are coerced to logical if possible." ``` --- ## More `if` examples ```r if (c(F, T, T)) { print("How many logical values can if handle?") } ``` ```r x <- c(1, 2, 3, 4) if (x %in% 3) { print("This works?") } ``` ```r if (c(1, 0, 1)) { print("Other types are coerced to logical if possible.") } ``` ``` #> [1] "Other types are coerced to logical if possible." ``` --- ## `if` is not vectorized To remedy this potential problem of a non-vectorized `if`, you can 1. try to collapse a logical vector of length greater than 1 to a logical vector of length 1 with functions - `any()` - `all()` 2. use a vectorized conditional function such as `ifelse()` or `dplyr::case_when()`. --- ## Functions `any()` and `all()` ```r x <- c(-5, 0, 5, 10, 15) any(x >= 5) ``` ``` #> [1] TRUE ``` ```r all(x >= 5) ``` ``` #> [1] FALSE ``` <br/><br/> Functions `any()` and `all()` require a logical vector as input. --- ## Vectorized `if` ```r z <- c(-4:-1, 1:3) z ``` ``` #> [1] -4 -3 -2 -1 1 2 3 ``` ```r ifelse(test = z < 0, yes = "neg", no = "pos") ``` ``` #> [1] "neg" "neg" "neg" "neg" "pos" "pos" "pos" ``` <br> -- ```r x <- rnorm(n = 4, mean = 0, sd = 1) x ``` ``` #> [1] -0.5738760 0.7314262 0.4771350 1.1926576 ``` ```r ifelse(test = abs(x) > 3, yes = "outlier", no = "no outlier") ``` ``` #> [1] "no outlier" "no outlier" "no outlier" "no outlier" ``` --- ## Nested conditionals .pull-left[ ```r if (condition_one) { ## ## Code to run ## } else if (condition_two) { ## ## Code to run ## } else { ## ## Code to run ## } ``` ] .pull-right[ ```r x = 0 if (x < 0) { "Negative" } else if (x > 0) { "Positive" } else { "Zero" } ``` ``` #> [1] "Zero" ``` ] --- class: inverse, middle, center # Error action --- ## Execute error action Functions `stop()` and `stopifnot()` execute an error action. These are useful if you want to validate inputs or function arguments. ```r x <- -1 if (x < 0) { stop("Negative numbers not allowed!") } ``` ``` #> Error in eval(expr, envir, enclos): Negative numbers not allowed! ``` <br> -- ```r x <- c(3, 9, 28) stopifnot(any(x >= 0), all(x %% 3 == 0)) ``` ``` #> Error: all(x%%3 == 0) is not TRUE ``` If any of the expressions in function `stopifnot()` are not `TRUE`, then function `stop()` is called and an error message is shown. --- ## Exercises 1. What does each of the following return? Run the code to check your answer. ```r if (1 == "1") "coercion works" else "no coercion " ifelse(5 > c(1, 10, 2), "hello", "olleh") ``` <br> 2. Consider two vectors, `x` and `y`, each of length one. Write a set of conditionals that satisfy the following. - If `x` is positive and `y` is negative or `y` is positive and `x` is negative, print "knits". - If `x` divided by `y` is positive, print "stink". - Stop execution if `x` or `y` are zero. Test your code with various `x` and `y` values. Where did you place the stop execution code? ??? ## Solutions 1. .solution[ ```r if (1 == "1") "coercion works" else "no coercion " ``` ``` #> [1] "coercion works" ``` ```r ifelse(5 > c(1, 10, 2), "hello", "olleh") ``` ``` #> [1] "hello" "olleh" "hello" ``` ] 2. .solution[ ```r x <- 4 y <- -10 if (x == 0 | y == 0) { stop("One of x or y is 0!") } else if (x / y > 0) { print("stink") } else { print("knits") } ``` ``` #> [1] "knits" ``` ] --- class: inverse, middle, center # Loops --- ## Loop types R supports three types of loops: `for`, `while`, and `repeat`. ```r for (item in vector) { ## ## Iterate this code ## } ``` ```r while (we_have_a_true_condition) { ## ## Iterate this code ## } ``` ```r repeat { ## ## Iterate this code ## } ``` In the `repeat` loop we will need a `break` statement to end iteration. --- ## `for` loop A `for` loop allows you to iterate code over items in a vector. ```r k = 0 for (i in c(2, 4, 6, 8)) { print(i ^ 2) k <- k + i ^ 2 } ``` ``` #> [1] 4 #> [1] 16 #> [1] 36 #> [1] 64 ``` ```r k ``` ``` #> [1] 120 ``` ```r for (i in c(2, 4, 6, 8)) { i ^ 2 } ``` *Automatic printing is turned off inside loops.* ??? .tiny[ ```r i <- 10 for (i in 1:3) { print("Today is Tuesday.") } ``` ``` #> [1] "Today is Tuesday." #> [1] "Today is Tuesday." #> [1] "Today is Tuesday." ``` ```r i ``` ``` #> [1] 3 ``` ] Variable `i` is overwritten when `for` is executed. Thus, `i` is assigned to the current environment. Convention is to use `i`, `j`, `k` as loop items. It will be best to avoid using these as named objects elsewhere in your code. --- ## `while` loop A `while` loop will iterate code until a given condition is `FALSE`. ```r i <- 1 res <- rep(0, 10) i ``` ``` #> [1] 1 ``` ```r res ``` ``` #> [1] 0 0 0 0 0 0 0 0 0 0 ``` ```r while (i <= 10) { res[i] <- i ^ 2 i <- i + 1 } res ``` ``` #> [1] 1 4 9 16 25 36 49 64 81 100 ``` --- ## `repeat` loop A `repeat` loop will iterate code until a `break` statement is executed. ```r i <- 1 res <- rep(NA, 10) repeat { res[i] <- i ^ 2 i <- i + 1 if (i > 10) {break} } res ``` ``` #> [1] 1 4 9 16 25 36 49 64 81 100 ``` --- ## Loop keywords: `next` and `break` - `next` exits the current iteration and advances the looping index - `break` exits the loop - Both `break` and `next` apply only to the innermost of nested loops. ```r for (i in 1:10) { * if (i %% 2 == 0) {next} print(paste("Number ", i, " is odd.")) * if (i %% 7 == 0) {break} } ``` -- ``` #> [1] "Number 1 is odd." #> [1] "Number 3 is odd." #> [1] "Number 5 is odd." #> [1] "Number 7 is odd." ``` --- ## Ancillary loop functions You may want to loop over indices of an object as opposed to the object's values. To do this, consider using one of `length()`, `seq()`, `seq_along()`, and `seq_len()`. .pull-left[ ```r 4:7 ``` ``` #> [1] 4 5 6 7 ``` ```r length(4:7) ``` ``` #> [1] 4 ``` ```r seq(4, 7) ``` ``` #> [1] 4 5 6 7 ``` ] .pull-right[ ```r seq_along(4:7) ``` ``` #> [1] 1 2 3 4 ``` ```r seq_len(length(4:7)) ``` ``` #> [1] 1 2 3 4 ``` ```r seq(4, 7, by = 2) ``` ``` #> [1] 4 6 ``` ] <br> Iterating over `seq_along(x)` is a better option than `1:length(x)`. --- ## Loop tips 1. Preallocate your output object when possible. 2. Don't use a `while` or `repeat` loop if a `for` loop is possible. 3. Don't use any type of loop if vectorization is possible. .pull-left[ Slow... ```r a <- c() for (i in seq_len(10)) { a <- c(a, i ^ 3) } ``` ] .pull-right[ Faster... ```r a <- numeric(10) for (i in seq_len(10)) { a[i] <- i ^ 3 } ``` ] Even faster... ```r (1:10) ^ 3 ``` --- ## Exercises 1. Consider the vector `x` below. ```r x <- c(3, 4, 12, 19, 23, 49, 100, 63, 70) ``` Write R code that prints the perfect squares in `x`. <br><br> 2. Consider `z <- c(-1, .5, 0, .5, 1)`. Write R code that prints the smallest non-negative integer `\(k\)` satisfying the inequality `$$\lvert cos(k) - z \rvert < 0.001$$` for each component of `z`. ??? ## Solution 1. .solution[ ```r x <- c(3, 4, 12, 19, 23, 49, 100, 63, 70) for (i in x) { if (sqrt(i) %% 1) { next } print(i) } ``` ``` #> [1] 4 #> [1] 49 #> [1] 100 ``` ] 2. .solution[ ```r for (z in c(-1, .5, 0, .5, 1)) { k <- 0 while (abs(cos(k) - z) >= .001) { k <- k + 1 } print(k) } ``` ``` #> [1] 22 #> [1] 21766 #> [1] 40459 #> [1] 21766 #> [1] 0 ``` ] --- ## References - Deborah Nolan & Duncan Temple Lang (2010) Computing in the Statistics Curricula, The American Statistician, 64:2, 97-107, DOI: 10.1198/tast.2010.09132 - Piatetsky, G. (2019). Python leads the 11 top Data Science, Machine Learning platforms: Trends and Analysis. Kdnuggets.com. Retrieved 21 August 2019, from https://www.kdnuggets.com/2019/05/poll-top-data-science-machine-learning-platforms.html - Which Programming Language Should Data Scientists Learn First?. (2019). Medium. https://towardsdatascience.com/which-programming-language-should-data-scientists-learn-first-aac4d3fd3038 - Grolemund, G., & Wickham, H. (2019). R for Data Science. https://r4ds.had.co.nz/ - Wickham, H. (2019). Advanced R. https://adv-r.hadley.nz/